Tracking subtle brain mutations, systematically: Tool can trace and spatially map 'mosaic' mutations in the brain

DNA sequences were once thought to be identical from cell to cell, but it's increasingly understood that mutations can arise during brain development that affect only certain groups of brain cells. A technique developed at Boston Children's Hospital allows these subtle mutation patterns to be traced and mapped spatially for the first time. This capability is a significant advance for genetics research and provides a new way to study both the normal brain and brain disorders such as epilepsy and autism.
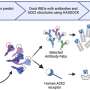
Described in the January 7th issue of Neuron, the technique uses "deep," highly sensitive whole-genome sequencing of single neurons and a new technology that identifies inserted bits of DNA caused by retrotransposons, one of several kinds of so-called somatic mutations that can arise as the brain develops.
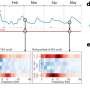
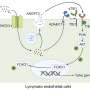
The technique picks up somatic mutations that affect just a fraction of the brain's cells, in a "mosaic" pattern. It also allows "lineage tracing," showing when during brain development the mutations arise and how they spread through brain tissue as the mutated cells grow, replicate and migrate, carrying the mutation with them.
"There is a lot of genetic diversity from one neuron to the other, and this work gets at how somatic mutations are distributed in the brain," says Christopher Walsh, MD, PhD, chief of Genetics and Genomics at Boston Children's and co-senior author on the paper. "Some mutations may occur on one side of the brain and not the other. Some may be 'clumped,' affecting just one gyrus [fold] of the brain, disrupting just a little part of the cortex at a time."
The study examined brain tissue from a deceased 17-year-old who had been neurologically normal, sampling in more than 30 brain locations. It builds on work published by the Walsh lab in 2012, which developed methods to sequence the genomes of single neurons, and represents the first time single neurons have been sequenced in their entirety. The single-cell technique is better at detecting subtle mosaicism than usual DNA sequencing methods, which sequence many thousands or millions of cells mixed together and read out an average for the sample.
Somatic brain mutations, affecting just pockets of cells, can be harmful, and have been suggested as a possible cause of neurodevelopmental disorders such as autism, epilepsy or intellectual disability (see this review article for further background). But they also can be completely benign or have just a subtle effect.
"Our findings are intriguing because they suggest that every normal brain may in fact be a mosaic patchwork of focal somatic mutations, though in normal individuals most are likely silent or harmless," says Gilad Evrony, PhD, in the Walsh Lab, co-first author on the Neuron paper. "These same technologies can now be used to study the brains of people who died from unexplained neuropsychiatric diseases to determine whether somatic mutations may be the cause."
Finally, says Evrony, the findings provide a proof-of-principle for a systematic way of studying how brain cells disperse and migrate during development, "something that has not been possible to do before in humans," he says.
Co-first author Alice Eunjung Lee, PhD, from the lab of Peter Park, PhD, at the Center for Biomedical Informatics at Harvard Medical School, developed the study's retrotransposon analysis tool, which detects somatic retrotransposon mutations in single-cell sequencing data.
Mirroring these findings, study published by Walsh's lab in 2014 used single-neuron sequencing to detect copy number variants—another type of mutation affecting the number of copies of chromosomes or chromosome fragments. The study found that these mutations can occur in both normal and neurologically diseased brains.