New mathematical model explains variability in mutation rates across the human genome

It turns out that the type, how frequent, and where new mutations occur in the human genome depends on which DNA building blocks are nearby, found researchers from the Perelman School of Medicine at the University of Pennsylvania in an advanced online study published this week in Nature Genetics.
"We developed a mathematical model to estimate the rates of mutation as a function of the nearby sequences of DNA 'letters'—called nucleotides—in the human genome," said senior author Benjamin F. Voight, PhD, an assistant professor in the department of Systems Pharmacology and Translational Therapeutics and the department of Genetics. "This new model not only provides clues into the process of mutation, but also helps discover possible genetic risk factors that influence complex human diseases, such as autism spectrum disorder."
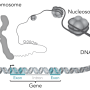
This study focuses on the probability that any given nucleotide in the human genome—one of the four letters (A, C, G or T for adenine, cytosine, guanine or thymine) of the DNA alphabet—is changed. Voight focused on the simplest type of mutation, a "point" mutation in which a single letter is changed in a given sequence. Most of these changes—often called single nucleotide polymorphisms (SNPs), or "snips"—are usually not harmful to the functioning of the human body. Nevertheless, Voight examined why some sequences are more prone to mutate, whereas others are not.
"The crux of the paper examines the dependency of mutation rate on which nucleotides are one, two, or three bases away from either side of a SNP," Voight said. "We already know about one situation in which this placement matters: DNA sequences in the genome where methyl groups are attached to the cytosine nucleotide, also known as CpG sites, are hotspots for mutation. But are there other types of local sequences that matter beyond these?"
To address this question, Voight and graduate student Varun Aggarwala, a doctoral candidate from the Genomics and Computational Biology graduate group, devised a mathematical model applicable to SNP data found in humans. Their approach took advantage of publicly available data from thousands of human subjects sampled from across the globe, namely from the 1000 Genomes Project. These individuals were sequenced as part of an international initiative to characterize the genetic variation that naturally occurs in human populations.
What they found was surprising: Knowing the three nucleotides flanking either side of a given SNP, for a total of seven nucleotides, predicted up to 93 percent of the variability in the chance of finding a SNP in a given sequence in individuals whose genome sequences are in the 1000 Genomes Project database. In addition, their model uncovered several distinctive sequences of local nucleotides that were not previously known to be prone to mutation.
"It turns out there are indeed DNA sequences beyond CpG sites that are also prone to mutation," Voight said. "What is not immediately obvious is why. The initial rates and our model need to be investigated more deeply to decipher the basic mechanisms that induce mutation in human genomes."
Another finding questioned the assumption that methylated CpG sites always have the same rate of mutation. "I think it is commonly assumed that all CpG sequences mutate at the same rate, though our results indicate far more variability that we expected," Voight said. Using another publicly available database that measured the methylation states at CpG sites across several individuals, Voight and Aggarwala found that the frequency in which different sequence contexts were methylated could not fully explain differences in mutation rates at these sites. "This certainly indicates the possibility of additional genetic mutation phenomenon at CpG hotspots that change how prone these sites are to mutate, for example how well DNA-repair machinery can correct new mutations that might arise," Voight said.
Beyond gleaning clues for different ways mutations occur, Voight and Aggarwala also examined applications of their model to human disease, providing ways to rank which newly discovered mutations identified from clinical genetics studies are the most likely to result in disease. Computational predictive measurements such as these are used to help prioritize rare or new gene variants discovered from these studies for follow-up investigation. Voight and Aggarwala focused on a set of autism sequencing studies by looking for genes with an excess of new mutations in children with autism not otherwise found in parents. When they applied their model to these data, they found an improvement over existing methods for predicting which rare or new mutations were associated with human disease.
"We were able to refine the focus somewhat on likely pathogenic variants for follow-up work, though we'll need quite a bit more work to correctly pinpoint the right variants and genes for autism or even Alzheimer's disease where sequencing data is readily available," Voight said.
He credits not only the large amount of publicly available data, but careful and dedicated efforts over an extended period as major contributing factors to be able to evaluate and refine their proposed mathematical model. "The exciting part of this work is not just what we've found, but the spectrum of new questions that we will be able to systematically address in the next few years. While building solid foundations takes time, the next set of scientific 'skyscrapers' built on these foundations will absolutely persist longer and reach higher as a result."
More information: Varun Aggarwala et al. An expanded sequence context model broadly explains variability in polymorphism levels across the human genome, Nature Genetics (2016). DOI: 10.1038/ng.3511