CRISPR study reveals unexpected roles of non-coding RNAs
UC San Francisco researchers have taken a major step toward understanding the function of the tens of thousands of human genes that do not code for proteins, a phenomenon considered one of the key remaining mysteries of the human genome. New findings , which focused on the roles of these genes in human cancer cells, suggest a possible new strategy for targeting the disease.
In recent years, researchers have recognized that non-coding regions of the genome—long dismissed as "junk DNA"—are actually key players in cell biology, development, and disease. However, the vast majority of these regions have not yet been extensively studied.
Now UCSF scientists have developed an approach to studying the function of genes that produce RNA transcripts but no protein – called long non-coding RNAs (lncRNAs)—at an unprecedented scale. In a proof of principle experiment, the researchers set out to look for common lncRNAs required for the growth of many different types of cancer cells. To their surprise, they found that each cancer cell line they tested relied upon a different set of IncRNAs for growth and survival, suggesting that these molecules might be promising targets for precision cancer therapies.
The new study—published online by the journal Science on Thursday, Dec. 15, 2016—was conducted jointly by the UCSF labs of Daniel Lim, MD, PhD, an associate professor of neurological surgery and member of the Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research at UCSF and Jonathan Weissman, PhD, a Howard Hughes Medical Investigator and professor of cellular and molecular pharmacology at UCSF.
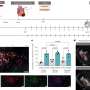
Researchers from the two labs teamed up to create a genome-scale platform to study lncRNA biology using CRISPR-based interference (CRISPRi), a technique developed in the Weissman lab in collaboration with Stanley Qi, PhD, now of Stanford University, that enables researchers to precisely tune the activity of specific genes up or down.
The UCSF team assembled a large library of CRISPR guide RNAs against 16,401 different lncRNA genes—a large number of those that are currently known—then used CRISPRi to selectively inactivate each gene in an array of seven human cell lines, including six cancer cell types and a line of induced pluripotent stem cells (iPSCs) supplied by co-author Bruce Conklin, MD, a professor of medicine at UCSF and the academically affiliated Gladstone Institute of Cardiovascular disease.
In collaboration with the lab of Howard Chang, MD, PhD, of Stanford University, the team found that although about 5,000 different lncRNA genes were actively expressed in each cell line, overall only 499 lncRNAs significantly impacted cellular growth when they were inactivated. Moreover, although the researchers had initially thought that there would be a set of lncRNAs that were essential to the survival of all cells, no such core set of essential lncRNAs were found. In fact, they found that the opposite was true: 89 percent of the "essential" lncRNAs were crucial for only one cell type, and had no effect on the others.
"This was a big surprise," said Lim, whose lab has been studying the role of lncRNAs in brain development. "Typically, protein-coding genes that are essential for one type of cell are also crucial for a broad range of cell types. But with lncRNAs we see an unexpected but exquisite specificity."
The researchers also examined how silencing particular lncRNAs impacted gene expression in the seven cell types. They found that when an lncRNA was "essential" to a particular cell type silencing it strongly modulated the activity of up to hundreds of other genes. In other cells, silencing the same lncRNA had no effect on the expression of any other genes.
"I think of it like the power-supply under your desk," Lim said. "If you've got your computer and a lot of peripherals plugged into it, tripping that switch can cause a very bad day. But if nothing's plugged in, you can flip the switch as much as you like and nothing happens."
Platform opens up study of non-coding RNA's role in evolution, development and disease
The seed of the new approach to studying lncRNAs came from conversations between the paper's two lead authors, John Liu and Max Horlbeck, who are MD-PhD students in the Lim and Weissman labs, respectively, and who also happened to be housemates at the outset of the project.
"The Lim lab had been spending a lot of time testing different lncRNAs one at a time to see if they were relevant to brain tumors," said Liu. "But that was incredibly slow. We had so few examples of lncRNA function, and it was very difficult to predict which lncRNAs would be important."
For their part, the Weissman lab had also grown interested in lncRNAs: "We had been working for several years to develop CRISPRi as a way of screening protein-coding genes," Horlbeck said. "But we realized that because the technique works by tuning down how much RNA a specific gene makes, it's a perfect tool to investigate the role of non-coding RNAs, which are hard to study with other methods at this scale."
The researchers say they are eager to broaden the study of lncRNA biology by sharing their techniques, which open up many fundamental questions about the biological function of lncRNAs in development and evolution, as well as their potential utility for precision medicine approaches to cancer treatment.
"One critical question is whether different lncRNAs are essential for the growth of cancerous cells in particular, but not healthy cells," Weissman said. "If so, they could potentially be targeted to impede tumors without being toxic to the rest of the body, and that would be very attractive for the pharmaceutical industry."
More information: S. J. Liu et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells, Science (2016). DOI: 10.1126/science.aah7111