Scientists identify unique cell-signaling system in some S. pneumoniae strains
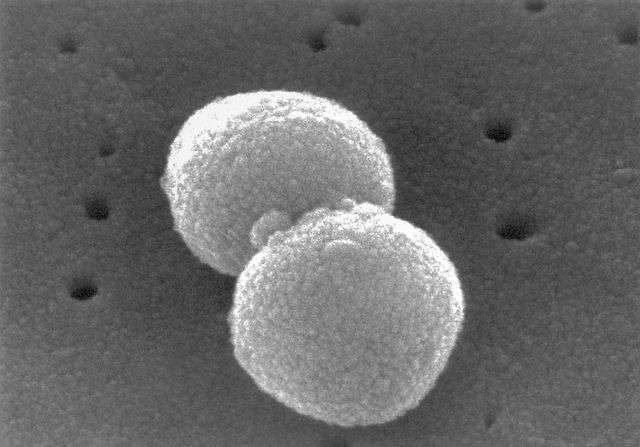
Some strains of Streptococcus pneumoniae—a disease-causing bacterium—possess a cell-to-cell signaling system that may influence gene expression and virulence in co-colonizing strains, according to a study published in PLOS Pathogens.
S. pneumoniae is widespread in the human respiratory tract, but usually does not cause any symptoms. However, in children and the elderly, it is a major cause of pneumonia, meningitis, and other potentially deadly diseases. Of particular interest is a group of S. pneumoniae strains known as PMEN1, which are pandemic and multi-drug resistant.
To better understand what makes PMEN1 strains so successful, Anagha Kadam of Carnegie Mellon University, Pennsylvania, and colleagues performed a genetic screen to identify genes possessed by PMEN1 strains, but not by other strains. This revealed that almost all PMEN1 strains have a unique genomic region consisting of several specific genes.
The team found that two of these genes, phrA2 and tprA2, interact to regulate S. pneumoniae gene expression. The protein encoded by tprA2 inhibits expression of phrA2 and other neighboring genes, including a gene known as lcpA. In mice, the researchers found, the TprA2 protein does not affect S. pneumoniae colonization in the nasopharynx, but it does protect against lung infection. The findings suggest that this is a result of TprA2's control over lcpA expression.
Meanwhile, the phrA2 protein product relieves the inhibitory effects of TprA2, increasing lcpA expression. phrA2 expression is higher when the density of cells in an S. pneumoniae colony is higher. Evidence suggests that cells in high-density populations secrete the phrA2 protein product, and this protein then signals other cells to express phrA2.
Beyond its role in this novel TprA2/PhrA2 system, the PhrA2 protein also appears to be able to activate a second, ancestral signaling system that is widespread among S. pneumoniae, including in non-PMEN1 strains. This suggests that PMEN1 strains can influence both their own gene expression and that of non-PMEN1 strains. This is highly relevant since multi-strain infections are very common.
Phylogenetic analysis suggests that the genes in the TprA2/PhrA2 system initially found their way into an ancestral PMEN1 strain through horizontal gene transfer, instead of being passed from a dividing cell to its daughter cells. The authors note that this may be the first example of a horizontally transferred regulatory system that has integrated with ancestral network to allow gene regulation across strains.
More information: Kadam A, Eutsey RA, Rosch J, Miao X, Longwell M, Xu W, et al. (2017) Promiscuous signaling by a regulatory system unique to the pandemic PMEN1 pneumococcal lineage. PLoS Pathog 13(5): e1006339. DOI: 10.1371/journal.ppat.1006339