Mutation found in patients without a nose
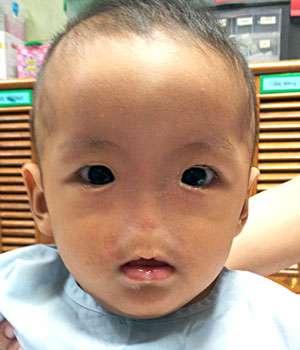
A mutated gene in patients lacking a nose has been identified by an international team, a first step toward understanding nose development and possible therapies for another condition.
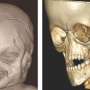
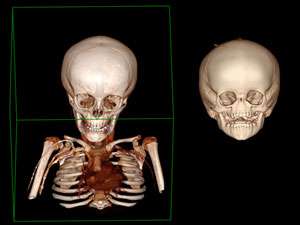
The team, led by researchers from Bruno Reversade's lab at the A*STAR Institute of Medical Biology and the Institute of Molecular and Cell Biology, sequenced DNA from 14 people with Bosma arhinia microphthalmia syndrome (BAMS). BAMS is a congenital condition in which patients are born without a nose, and often with eye defects and stunted release of sexual hormones.
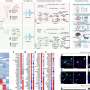
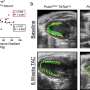
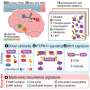
All 14 people had mutations in the gene SMCHD1, none of which were present in the parental DNA samples, indicating that they were spontaneous mutations. SMCHD1 is known to regulate chemical modifications to DNA which silence or repress genes, and this study showed that it is expressed in the developing nasal and optical tissues of mouse embryos.
SMCHD1 is already implicated in a more common disease, facioscapulohumeral muscular dystrophy (FSHD). "FSHD is a very different disease," says Shifeng Xue, a research fellow in the Reversade lab who led the project. "It's an adult onset degenerative disease, whereas BAMS is an embryological developmental disease. The only similarity is that they have mutations in the same gene."
The diseases are also driven by entirely different types of mutations. While FSHD2 results from mutations that knock out SMCHD1, the team believes that BAMS is caused by increased activity of the gene. Introducing the mutant gene into frog embryos led to craniofacial defects similar to those of BAMS patients, and the team observed the same defects when they increased expression of the unmutated form of the gene.
While SMCHD1 is clearly a key gene regulating nose development, its precise function remains a mystery. "SMCHD1 is a repressor," explains Xue. "We think the gain-of-function mutations cause it to silence more genes than it's supposed to, including genes involved in nose development."
"Finding SMCHD1 gives us a handle to tackle this question," she adds. Developmental genetics research often uses mice as models, but there are no mutant mice lacking a nose; mice cannot mouth breathe, so they die if they are born without a nose. "Now that we know this gene is a master regulator, we can explore its downstream targets and signaling pathways," says Xue.
Reversade calls the discovery "a fascinating example of how rare conditions can provide insights into common diseases," adding that the team hopes to translate their findings into potential FSHD therapies.
More information: Christopher T Gordon et al. De novo mutations in SMCHD1 cause Bosma arhinia microphthalmia syndrome and abrogate nasal development, Nature Genetics (2017). DOI: 10.1038/ng.3765