Editing false positives from cancer dependency maps drawn with CRISPR
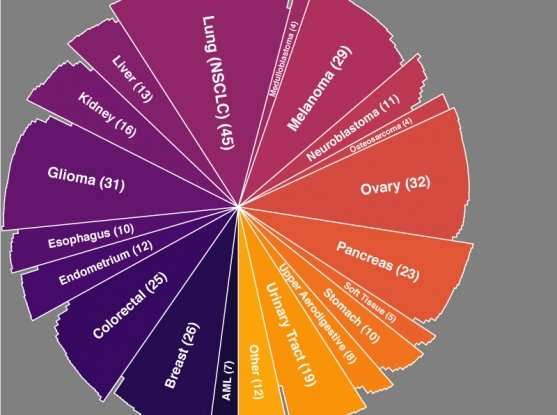
The Broad Cancer Dependency Map team adds CRISPR-based data from 342 cancer cell lines to their growing catalog of genetic dependencies in cancer, and a new method for ensuring that data's accuracy.
Genome-scale CRISPR-based knockout screens are powerful tools for pinpointing cells' genetic dependencies—that is, genes that cells require for their survival and/or proliferation. However, such CRISPR screens are sensitive to a phenomenon called the copy number effect, where genes that have been repeatedly duplicated within a cell (as commonly happens in cancer cells) can be flagged as essential regardless of whether they are or not.
To limit such false-positive hits, the Broad Institute's Cancer Dependency Map project—a joint effort bringing together researchers from the Broad Cancer Program's Project Achilles and Cancer Data Science teams, the institute's Genetic Perturbation Platform, and other Broad groups—has developed CERES, a computational method that corrects pooled CRISPR screen data for the copy number effect and provides an unbiased view of cancer cells' genetic dependencies.
As they revealed in Nature Genetics, the team benchmarked CERES against genome-scale CRISPR-Cas9 data from 342 cancer cell lines (the largest CRISPR knockout dataset generated in cancer lines to date) curated by the Broad-Novartis Cancer Cell Line Encyclopedia (CCLE). The method greatly reduced false-positive readouts in the data, pinpointing known dependencies (e.g., KRAS mutations) and allowing new dependencies to become apparent.
The new dependency data complement the Dependency Map team's ongoing efforts to use functional genomic technologies like CRISPR and RNA interference (RNAi) to locate vulnerabilities that arise within cancer cells as they compensate for the loss of critical genes due to mutations or expression changes. Earlier this year the team announced that they had cataloged 769 strong genetic dependencies across 501 CCLE-curated cell lines using genome-scale RNAi screens—the fruits of a nearly 10-year effort.
CERES joins two prior computational methods the team has developed to filter false-positive results from functional genomic screen data: ATARiS and DEMETER, both of which weed out so-called seed effects that commonly plague RNAi data.
More information: Robin M Meyers et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells, Nature Genetics (2017). DOI: 10.1038/ng.3984