Study finds gut microbiome can control antitumor immune function in liver
Scientists have found a connection between bacteria in the gut and antitumor immune responses in the liver. Their study, published May 25 in Science, was led by researchers in the Center for Cancer Research (CCR) at the National Cancer Institute (NCI). It showed that bacteria found in the gut of mice affect the liver's antitumor immune function. The findings have implications for understanding the mechanisms that lead to liver cancer and for therapeutic approaches to treat them. NCI is part of the National Institutes of Health.
"What we found using different tumor models is that if you treat mice with antibiotics and thereby deplete certain bacteria, you can change the composition of immune cells of the liver, affecting tumor growth in the liver," said Tim Greten, M.D., of NCI's CCR, who led the study. "This is a great example of how what we learn from basic research can give us insight into cancer and possible treatments."
The microbiome is the collection of bacteria and other microorganisms that live in or on the body. In humans, the greatest proportion of the body's total microbiome is in the gut. Despite extensive research into the relationship between the gut microbiome and cancer, the role of gut bacteria in the formation of liver cancer has remained poorly understood.
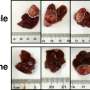
To investigate whether gut bacteria affect the development of tumors in the liver, Dr. Greten and his team carried out a series of experiments with mice. They used three mouse liver cancer models, and found that when they depleted gut bacteria using an antibiotic "cocktail," the mice that had the antibiotics developed fewer and smaller liver tumors and had reduced metastasis to the liver.
The investigators next studied the immune cells in the liver to understand how the depletion of gut bacteria suppressed tumor growth in the liver of the antibiotic-treated mice. Antibiotic treatment increased the numbers of a type of immune cell called NKT cells in the livers of the mice. Further experiments showed that, in all three mouse models, the reduction in liver tumor growth that resulted from antibiotic treatment was dependent on these NKT cells. Next, they found that the accumulation of the NKT cells in the liver resulted from an increase in the expression of a protein called CXCL16 on cells that line the inside of capillaries in the liver.
"We asked ourselves, why do mice treated with antibiotics have more CXCL16 production in these endothelial cells?" Dr. Greten said. "That was the critical point, when we found that bile acids can control the expression of CXCL16. We then did further studies, and found that if we treat mice with bile acids, we can actually change the number of NKT cells in the liver, and thereby the number of tumors in the liver."
Bile acids are formed in the liver and help break down fats during digestion.
Finally, the investigators found that one bacterial species, Clostridium scindens, controls metabolism of bile acids in the mouse gut—and ultimately CXCL16 expression, NKT cell accumulation, and tumor growth in the liver.
Dr. Greten explained that while many studies have shown an association between gut bacteria and immune response, this study is significant in that it identifies not just a correlation, but a complete mechanism of how bacteria affect the immune response in liver. In the same study, the researchers found that bile acids also control the expression of the CXCL16 protein in the liver of humans and wrote that, though these results are preliminary, the novel mechanism described in this study could potentially apply to cancer patients.
This press release describes a basic research finding. Basic research increases our understanding of human behavior and biology, which is foundational to advancing new and better ways to prevent, diagnose, and treat disease. Science is an unpredictable and incremental process—each research advance builds on past discoveries, often in unexpected ways. Most clinical advances would not be possible without the knowledge of fundamental basic research.
More information: C. Ma el al., "Gut microbiome–mediated bile acid metabolism regulates liver cancer via NKT cells," Science (2018). science.sciencemag.org/cgi/doi … 1126/science.aan5931