Analyzing single-cell landscapes
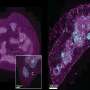
Single-cell RNA sequencing is a powerful tool for studying cellular diversity, for example in cancer where varied tumor cell types determine diagnosis, prognosis and response to therapy. Single-cell technologies generate hundreds to thousands of data points per sample, generating a need for new methods to define cell populations across different single-cell landscapes.
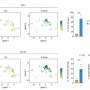
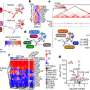
Qi Liu, Ph.D., Ken Lau, Ph.D., and colleagues have developed a new tool, sc-UniFrac, to quantify diverse cell types in single-cell studies. The tool compares hierarchical trees that represent single-cell landscapes and allows cells that drive differences to be identified as unbalanced branches on the trees.
Reporting in PLOS Biology, the investigators demonstrated the utility of sc-UniFrac in multiple applications, including regional specification of brain cells and identification of altered cells in tumor samples. The authors expect that sc-UniFrac will facilitate single-cell studies, in particular studies aimed at tracking how tumor cell populations evolve during disease progression and respond to drug treatments.
More information: Qi Liu et al. Quantitative assessment of cell population diversity in single-cell landscapes, PLOS Biology (2018). DOI: 10.1371/journal.pbio.2006687