From imaging neurons to measuring their true activity
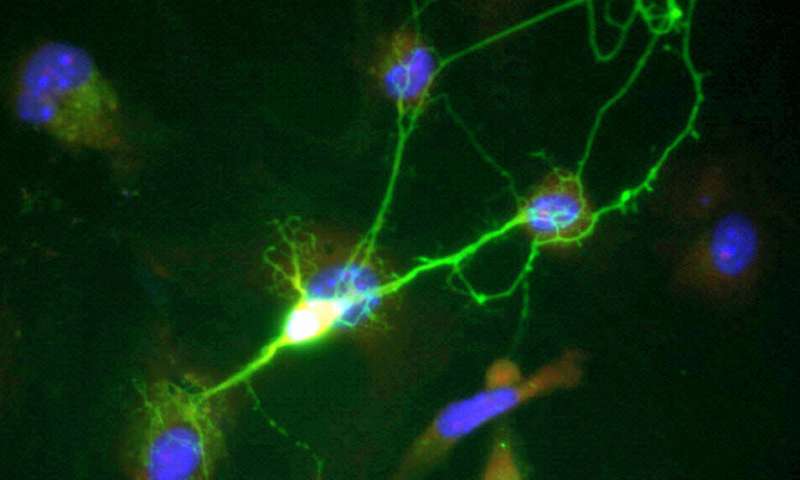
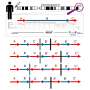
When neurons communicate with each another, they transmit—or "fire"—small electrical impulses called action potentials or spikes. These action potentials are the fundamental units of information processing in the brain. Today, neuronal activity is often measured by calcium imaging, which uses advanced microscopy to detect changes in the fluorescence of a calcium indicator inside neurons. This approach has become very popular because it can detect neuronal activity simultaneously in many neurons in the intact brain. However, rather than detecting the action potentials directly, it is an indirect measure of neuronal activity: The fluorescence signals depend on calcium influx through calcium channels in the cell membrane, which are activated by action potentials. Individual action potentials cause a transient increase and subsequent decrease in intracellular calcium concentration that is much slower than the action potential itself. The calcium signal measured by microscopy is therefore a slow, distorted and noisy "shadow" of the real electrical activity of a neuron. It is thus desired to reconstruct the true fluctuations in action potential firing rate from the measured calcium signals, which is no trivial task.
The relationship between calcium signals and spike rates is ideally assessed by simultaneous electrophysiological recordings and optical imaging of the calcium indicator signal in the same neuron. Such dual recordings can serve as "ground truth" to calibrate and optimize algorithms for the inference of spike rates from other calcium imaging data. Based on such ground truth datasets, various algorithms for spike inference have been developed, but they are generally complicated to use, with uncertain accuracy.
A history in reconstructing action potentials
FMI group leader Rainer Friedrich was among the first to develop an approach to reconstruct spikes from calcium imaging in this 2006 paper. The interest of Friedrich in the topic was then reactivated when his Ph.D. student Peter Rupprecht came up with the idea of using advanced machine learning to reconstruct spikes. After winning the "Spikefinder" challenge in 2017, Rupprecht dedicated his last months at the FMI to simultaneously measure calcium signals and action potentials from many different neurons and develop a new algorithm to infer spikes from calcium imaging. He then continued to work on this project and finalized it as a postdoc at the Brain Research Institute at the University of Zurich. Rupprecht's simple and effective algorithm, featured today in Nature Neuroscience, clearly outperformed all existing algorithms.
First Rupprecht compiled a large and diverse ground truth database from publicly available and newly performed recordings—using various species (mouse and zebrafish), types of neurons, calcium indicators, etc. He then developed a novel algorithm for spike inference that takes advantage of the ground truth dataset and is based on machine learning. By training the algorithm on the various datasets, he managed to create a kind of "universal" model that makes accurate predictions even for datasets that the algorithm has never been exposed to before.
A new standard
"There are two main reasons why Peter's approach turned out to be so successful," says Friedrich. "First he managed to get a very wide and diverse ground truth dataset. And then he had this brilliant idea of training the algorithm also on low quality data that included a lot of noise, which is the much more realistic scenario."
Friedrich expects this tool to become the new standard for neurobiologists to measure neuronal activity with calcium imaging, highlighting another advantage of the algorithm: the ease of use. "You perform your experiments, you upload your data in the tool, and in less than half an hour you get the results; no need to tweak any parameter, this is all done by the system." Moreover, the toolkit provides a platform to quantify the performance of other algorithms and to further improve action potential inference in the future, Friedrich says.
But is this new tool really going to make a difference for neurobiology experiments? "Of course," exclaims Friedrich. "The activity in the brain has a lot of structure on a timescale of a hundred milliseconds. With the much slower calcium signaling, this fine temporal structure is lost." The group leader then shares as an example an experiment where the new algorithm was applied in his own lab: The researchers measured the activity of 1,500 neurons simultaneously in the zebrafish brain during odor stimulation. They saw that the activity was changing over time but they only had a resolution of one or two seconds. With the new tool, they managed to push the temporal resolution by an order of magnitude. "Now we can see many things that we were unable to resolve before, such as ensembles of neurons that are activated in an orderly sequence when specific information is processed."
More information: Peter Rupprecht et al, A database and deep learning toolbox for noise-optimized, generalized spike inference from calcium imaging, Nature Neuroscience (2021). DOI: 10.1038/s41593-021-00895-5
The algorithm can be accessed here: github.com/HelmchenLabSoftware/Cascade