Genes associated with hearing loss visualised in new study
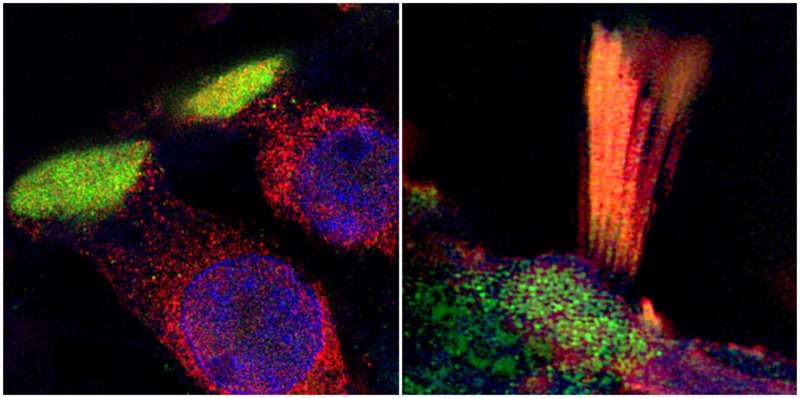
Researchers from Uppsala University have been able to document and visualize hearing loss-associated genes in the human inner ear, in a unique collaboration study between otosurgeons and geneticists. The findings illustrate that discrete subcellular structures in the human organ of hearing, the cochlea, are involved in the variation of risk of age-related hearing loss in the population. The study is published in BMC Medicine.
Hearing loss is a potentially debilitating condition that affects more than 1.23 billion people worldwide. The most common form of hearing loss, which represents 90% of all cases, is related to the degenerative effects of aging on hearing, i.e., age-related hearing loss or presbycusis. However, the molecular mechanisms that underlie the development of age-related hearing loss and individual variation in risk are poorly elucidated.
In the current study, a unique collaboration was established between otologists and geneticists at Uppsala University, which allowed for functional follow-up studies of candidate genes from genome-wide association studies (GWAS) using immunohistochemistry in the human cochlea.
"The cochlea, and in particular the hearing organ, the organ of Corti, is a highly vulnerable structure that is difficult to analyze since it is surrounded by the hardest bone in the body," says Helge Rask-Andersen, MD and Senior Professor at the Department of Surgical Sciences. "We have been able to study some of the molecular components of human hearing that are critical for the conversion of sound to nerve electric impulses."
Genetic variants at 67 genomic regions were found to contribute to increased risk of age-related hearing loss. Genome-wide association studies (GWAS) on hearing-related traits were performed in the UK Biobank, which has half a million participants from the United Kingdom. Genetic associations are difficult to interpret by themselves and follow-up experiments are often required before causal genes can be inferred.
"It is an amazing opportunity to be able to follow up our findings in human cochlear samples, since there are molecular differences between the hearing organ of humans and other mammals," says Mathias Rask-Andersen, Associate Professor at the Department of Immunology, Genetics and Pathology.
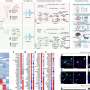
Candidate proteins from GWAS were visualized with immunofluorescent antibodies and super-resolution structured illumination microscopy (SR-SIM) by Dr. Wei Liu, MD and Associate Professor at the Department of Surgical Sciences. Several proteins were observed within the spiral ganglion, which contains the neuronal cell bodies that innervate the hair cells in the organ of Corti and carry neuronal impulses to the brain via the cochlear nerve.
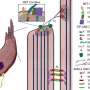
The researchers could also visualize hearing loss-associated proteins in discrete subcellular domains in the hair cells for the first time in humans, such as TRIO and F-actin-binding protein (TRIOBP) in the hair tufts (stereocilia) and LIM domain only protein 7 (LMO7) in the cuticular plate, which is an actin-rich structure that anchors stereocilia to the cell body. The stereocilia are the microscopic or nano-sized 'hairs' that protrude from the hair cells of the organ of Corti. They respond to mechanical vibrations from sounds that reach us and are transferred and amplified from the ear drum to the inner ear by the small middle ear bones.
Taken together, the findings from the current study demonstrate that common genetic variations associated with age-related hearing loss affect the structures of the cochlea, in particular the neuronal processes of the spiral ganglion, but also structures directly involved in the transduction of mechanical stimuli to neuronal impulses. This knowledge may help to better understand the biological mechanisms that lead to age-related hearing loss and generate strategies for prevention such as novel pharmacological treatments.
More information: Wei Liu et al, A combined genome-wide association and molecular study of age-related hearing loss in H. sapiens, BMC Medicine (2021). DOI: 10.1186/s12916-021-02169-0