SARS-CoV-2 hijacks antiviral human proteins to enter human cells
SARS-CoV-2 depends on the broadly antiviral interferon-induced human transmembrane proteins (IFITMs), to enter human cells and replicate inside them, according to research published this week in the Journal of Virology, a publication of the American Society for Microbiology.
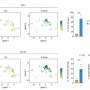
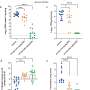
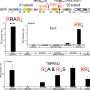
The investigators found that all 5 SARS-CoV-2 variants of concern—Alpha through Delta, and Omicron—"remain strongly dependent on antiviral transmembrane proteins, especially IFITM2," to replicate efficiently and to produce infectious progeny viruses, said Frank Kirchhoff, Ph.D., professor of virology, Ulm University Medical Center, Ulm, Germany.
"In addition, we show that an antibody against IFITM2 can protect human lung cells from SARS-CoV-2 infection," said Kirchhoff. "Our results suggest that IFITM2 may represent a highly unexpected target for a host-directed therapeutic approach. Targeting cellular factors in the host, rather than viral factors, reduces the risk of emergence of viral resistance."
The research aimed to find cellular factors that influence SARS-CoV-2 infection and to gain insight into innate immune defense mechanisms, as well as determinants of viral spread and pathogenesis. (Innate immunity is the body's first line of defense—detecting invaders such as viruses, bacteria, parasites and toxins, and then activating antiviral factors and immune cells to attack and destroy these invaders.)
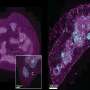
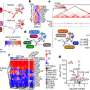
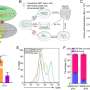
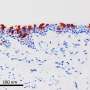
The researchers performed SARS-CoV-2 infection studies in a human epithelial lung cancer cell line expressing normal and experimentally reduced levels of IFITM proteins, then measured viral replication by quantifying viral RNA and infectious virus production. In addition, the team treated human lung cells with antibodies targeting IFITM2 or the viral ACE2 receptor [the major factor used by SARS-CoV-2 to gain entry into the cell] and found that both measures inhibit SARS-CoV-2 infection.
The team identified IFITM proteins as important interferon-inducible enhancers of SARS-CoV-2 infection that are expressed by all relevant target cells analyzed. "This has important implications for our understanding of the spread and pathogenesis of SARS-CoV-2. Additionally, our results provide insight into how SARS-CoV-2 avoids, or in this case, even exploits innate cellular defense mechanisms," said Kirchhoff.
As an expert on HIV, Kirchhoff's original approach was to determine whether cellular factors that restrict HIV might be active against SARS-CoV-2. "The eureka moment was when we realized that artificial overexpression of IFITMs inhibits SARS-CoV-2 as expected but—in striking and totally unexpected contrast—endogenous IFITMs in human lung cells were essential for efficient viral entry and replication," he said.
More information: Rayhane Nchioua et al, SARS-CoV-2 Variants of Concern Hijack IFITM2 for Efficient Replication in Human Lung Cells, Journal of Virology (2022). DOI: 10.1128/jvi.00594-22