March 31, 2023 report
This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
A cancer-wide analysis finds cancer-wide targets for tumor reduction
Researchers at the Washington University School of Medicine, St. Louis, have discovered a potential new target for cancer immunotherapy in transposable elements (TEs), short segments of DNA that can move around the genome.
In the paper, "Pan-cancer analysis identifies tumor-specific antigens derived from transposable elements" published in Nature Genetics, researchers used the Cancer Genome Atlas (TCGA), a database of more than 20,000 cancer samples covering 33 distinct cancer types and and 675 cancer cell lines.
The paper focuses on transposable elements (TEs) that exist in the human genome but are often dormant. These are, as the name implies, segments of DNA that can insert themselves into other genes. The classic example is a retrovirus that incorporates some of its genetic code in the genome of a host. The genome is prepared for this sort of intrusion, and continued replication and insertion is typically "turned off."
When cancers develop, they do so without much of the normal genetic failsafes in place, as they often originate from cells that have damaged DNA to begin with. In replicating, cancer allows these dormant TEs to regain their transposable ability within cancer cells and, according to the study, "drive expression of atypical transcripts that, at times, can be both highly expressed and widely shared across tumor samples."
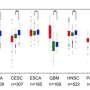
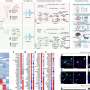
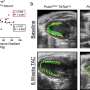
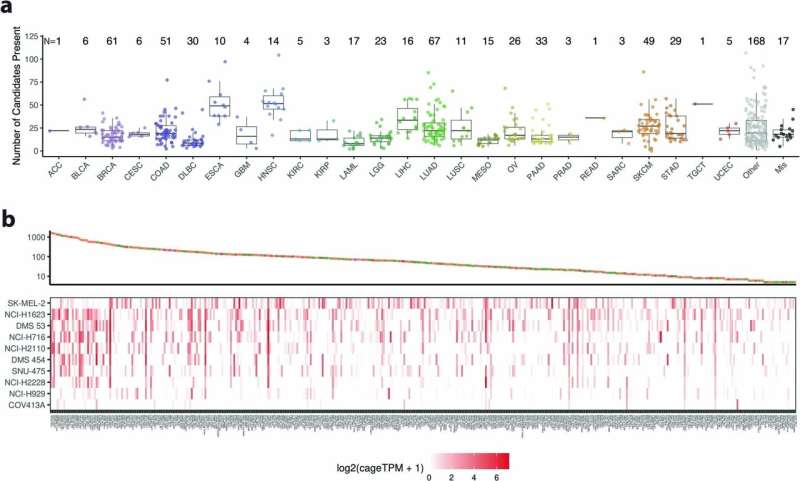
Because of this, the researchers were able to find TEs incorporated throughout 98% of tumors, across the distinct types, and found that "these transcripts have the potential to produce new peptides, which are not present in benign tissues," suggesting a novel way of targeting tumors, across cancer types.
The study identified 1,068 tumor specific TE candidates with the potential to generate shared tumor-specific TE-chimeric antigens (TS-TEAs). There are currently cancer treatments that target specific antigen types, and more research is needed to confirm that this pathway can be exploited to attack tumor cells with these TS-TEAs.
Medical science always operates in small, iterative steps of discovery, leading to caveat-filled potential pathways to conceiving of a cure. The difference here is that many of those pathways to a cure have already been traversed.
What the current study has uncovered is a mechanism within tumors that is creating possible targets for existing treatment approaches. This discovery could lead to new therapeutic approaches for cancer treatment, and there is a strong suggestion that with further research a single treatment type could developed to tackle multiple cancer types throughout the body.
More information: Nakul M. Shah et al, Pan-cancer analysis identifies tumor-specific antigens derived from transposable elements, Nature Genetics (2023). DOI: 10.1038/s41588-023-01349-3
© 2023 Science X Network