This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New cancer therapy target stops tumor cells from sharing resources
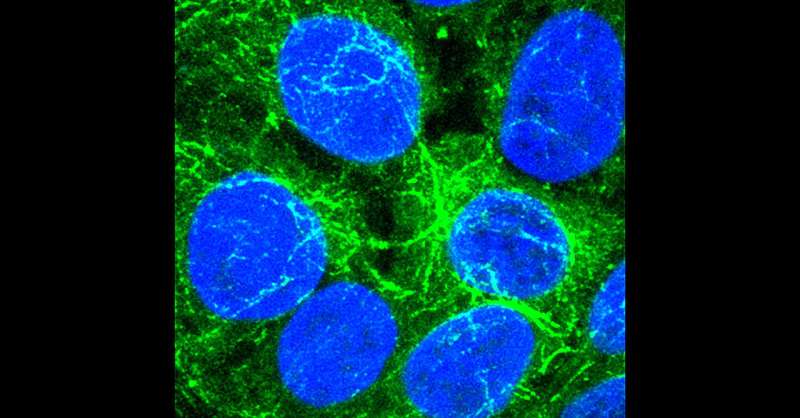
Researchers at University of California San Diego have discovered a process in which liver cells share molecules via vesicle exchange in order to multiply under conditions that would ordinarily suppress cell proliferation. They also found evidence that this process occurs in various types of cancer cells, paving the way for a new approach to tackling treatment resistance in cancer. The findings were published on October 17, 2023 in eLife.
"Understanding cell proliferation is a fundamental issue in both cancer research and biomedical science as a whole," said Gen-Sheng Feng, Ph.D., a professor of pathology at UC San Diego School of Medicine and of molecular biology at UC San Diego School of Biological Sciences. "While we studied the liver here because of its incredible regenerative potential, we've also found these vesicles in other types of cancer cells. We believe that this could be a universal mechanism that drives resistance to cancer therapies that target cell proliferation."
The body has many molecular processes in place to help control cell proliferation, turning it on and off as necessary in different circumstances and in different organs. This is well-exemplified by the liver, which has a high capacity for regeneration to support its important detoxification and metabolic functions.
"Cells in the liver multiply more quickly and effectively than any other cells in the body, which makes the liver the ideal place to study the biological processes that control cell division," said Feng. "These are the same processes that go awry in cancer, and so one promising approach to treating cancer is targeting cell proliferation."
In previous research, Feng and his team observed that a minority of liver cells in mice could still proliferate even when the cells were genetically engineered to lack a critical signaling enzyme required for cell proliferation. This enzyme, called Shp2, helps liver cells know when it's time to divide during liver regeneration. Shp2 is also a known target for treating various cancers, and Shp2 inhibitors are involved in several ongoing clinical trials.
The researchers discovered that liver cells have a clever way of compensating for the missing enzyme. By clustering together and exchanging tiny, protein-wrapped packages called vesicles, liver cells are able to communicate and share the biochemical materials needed to multiply, allowing them to proliferate even without Shp2.
"We've known for a long time that liver cells have other ways of regenerating even when lacking Shp2, and this study shows us how they do it," said Feng.
While the researchers made the discovery in noncancerous liver cells, they also uncovered evidence that cancer cells may use the same strategy to resist treatment and continue to divide. The vesicles that liver cells use to share molecules are marked by a protein called CD133, and the researchers also detected abnormally high amounts of this protein in a variety of human cancer cells.
"We believe we've discovered an important strategy that tumors use to resist therapy," said Feng. "This discovery could prove to be a powerful new target for cancer treatment, particularly as part of combination therapies to mitigate treatment resistance."
While the findings could lead to the discovery of new cancer therapies, they also suggest a new way of thinking about cancer initiation, progression and recurrence. CD133 is widely viewed as a marker of cancer stem cells, groups of cancer cells that are thought to be responsible for initiating tumors.
"Our findings could tell us that a cancer cell's 'stemness,' or ability to initiate and reinitiate tumors, isn't a central part of its identity, but a temporary state that can be induced or possibly even turned off," said Feng. "This could be a brand-new way of looking at tumor recurrence that could open doors for new treatments."
More information: Kota Kaneko et al, Identification of CD133+ intercellsomes in intercellular communication to offset intracellular signal deficit, eLife (2023). DOI: 10.7554/eLife.86824