This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
How a timekeeping gene affects tumor growth depends entirely on context
A new study has found that the circadian clock—which synchronizes physiological and cellular activities with the day-night cycle and is generally thought to be tumor suppressive—in fact has a contextually variable role in cancer.
"A lot of evidence suggests that the biological clock is broken in cancer cells, so we expected its disruption would fuel tumor growth in mouse models of melanoma," said Chi Van Dang, scientific director of the Ludwig Institute for Cancer Research, who led the study with Research Associate Xue Zhang. "But, contrary to our expectations, we found that loss of a gene known as Bmal1—a master regulator of the cellular clock—instead dampens the growth of melanoma tumors in mice."
The researchers' exploration of why this is the case, described in the current issue of Nature Communications, has uncovered previously unknown mechanisms of tumor suppression and resistance to immunotherapy in melanoma.
"Our findings may have implications for the design of chronotherapies, which seek to coordinate cancer treatments with biological rhythms to improve their efficacy and mitigate their side effects," said Zhang.
The circadian clock synchronizes the physiology of the body and the metabolism of each of its individual cells with the solar cycle, alternating phases of vibrant activity with periods of rest and recuperation. In healthy cells the intensity of metabolic activity oscillates with the biological clock. Cancer cells, however, rewire their metabolism to ensure a steady supply of the raw materials required for unrestrained proliferation. Since the periods of rest enforced by the circadian rhythms can undermine that agenda, they also tend to break their biological clocks.
The Dang lab, which has made seminal contributions to our understanding of cancer cell metabolism, has also explored links between the metabolic rewiring of cancer cells and their dysfunctional clocks. Most notably, studies led by Dang have shown that oncogenic versions of the Myc gene, a master regulator of cancer metabolism, suppress Bmal1 to disable the cell's metabolic brakes and fuel proliferation.
The slow growth of melanoma tumors lacking Bmal1 was not consistent with that finding. Looking for an explanation, the researchers examined how gene expression changed in these cells and saw that the loss of Bmal1 compromised the activity of another master regulator of gene expression, HIF1, that helps cancer cells adapt to low-oxygen conditions prevalent in many solid tumors. They also showed that expressing HIF1 at high levels in melanoma cells that lack Bmal1 restores the ability of those cells to generate tumors.
But would Bmal1 be tumor suppressive if expressed at sufficiently high levels? Apparently not, and it turns out that this has nothing to do with Bmal1's primary role as a regulator of gene expression. A version of the protein that could do everything ordinary Bmal1 does except bind to DNA—which it must do to regulate circadian gene expression—was an even better driver of tumor growth.
"That finding was a bit of a head-scratcher," said Dang. "It didn't fit with anything that we would expect."
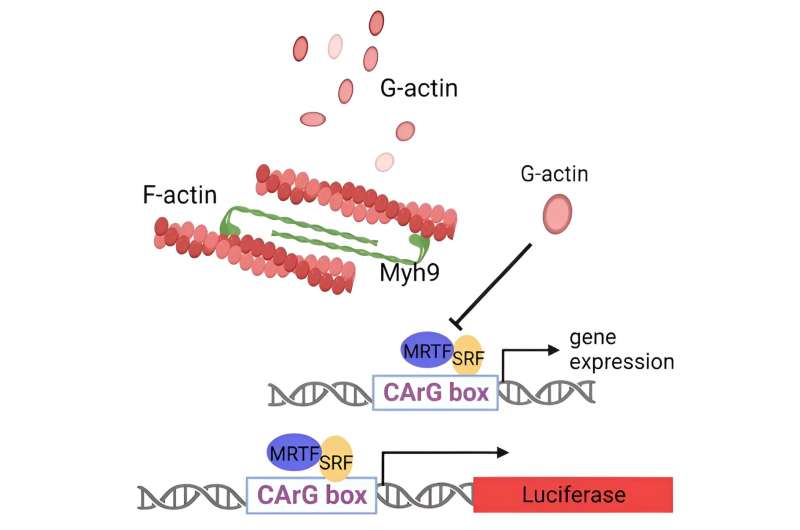
To solve the puzzle, the researchers conducted a sweeping analysis of protein interactions in Bmal1-overexpressing cells. They found that Bmal1 binds and sequesters a protein involved in cell motility, MYH9, that is thought to be a tumor suppressor—though how it exerts that effect was, until now, unclear. Aside from identifying the mechanism by which MYH9 suppresses tumorigenesis, the researchers show that deleting the MYH9 gene in melanoma cells boosts their ability to form tumors in mice.
Bmal1's sequestration of MYH9 induces several changes in the molecular signaling networks of cells. Its net effect, however, is to revert melanoma cells into a developmentally immature "mesenchymal" state that is seen in human tumors and is associated with resistance to immunotherapy. Indeed, the researchers show that melanoma cells expressing high levels Bmal1 form immunotherapy-resistant tumors in the mice.
With the support of Ludwig Harvard's Sandro Santagata and Peter Sorger, who lead the Ludwig Tumor Atlas project, the researchers employed advanced microscopy to show that tumors over-expressing Bmal1 bear molecular markers of the mesenchymal state. They also demonstrated that these tumors in mice resist immunotherapy because they are insufficiently infiltrated with immune cells that target cancer cells and tend to house immunosuppressive cells instead.
"Our findings overall show that Bmal1 drives tumorigenesis and therapeutic resistance in melanoma, suggesting that the effect of this regulatory protein on tumorigenesis most likely varies between cancer types," said Dang. "They may also help explain a recent discovery that prostate tumors expressing Bmal1 at high levels resist androgen therapy—though that will have to be confirmed with further research."
Beyond that, the findings have opened new avenues for research on mechanisms of tumor plasticity and cancer drug resistance that the Dang lab will now pursue.
More information: Xue Zhang et al, Cell state dependent effects of Bmal1 on melanoma immunity and tumorigenicity, Nature Communications (2024). DOI: 10.1038/s41467-024-44778-2