A striking link is found between the Fragile-X gene and mutations that cause autism
A team led by scientists at Cold Spring Harbor Laboratory (CSHL) publishes research today indicating a striking association between genes found disrupted in children with autism and genes that are targets of FMRP, the protein generated by the gene FMR1, whose dysfunction causes Fragile-X syndrome. The new study appears online April 25 in the journal Neuron.
Fragile-X syndrome is the most common cause of inherited intellectual disability. It is also counted among the autism spectrum disorders (ASDs) owing to the co-occurrence of autism-like symptoms in patients. A usually devastating disorder, Fragile X occurs when the FRM1 gene fails to direct nerve cells to manufacture FMRP, the FMR1-encoded protein, which plays a vital role in neural development and synaptic plasticity.
"A surprising proportion -- up to 20% -- of the candidate genes we see in our sample drawn from 343 autism families appear to be regulated by FMRP," says CSHL Research Investigator Dr. Michael Ronemus, co-first author of the new study. "Because of research connecting FMRP to the phenomenon of neuroplasticity, our work indicates a possible convergence of mechanisms causing autism," adds CSHL Professor Michael Wigler, the senior author of the study. Neuroplasticity is the process by which our brains become sensitized and desensitized to repetitive inputs.
Besides team leader Wigler, a geneticist, others who worked on the study included CSHL sequencing authority Dr. W. Richard McCombie, a sequencing team at Washington University, St. Louis, and Drs. Michael Schatz, Ivan Iossifov and Dan Levy of CSHL, all computational biologists.
Families studied by the team were part of the Simons Simplex Collection. This collection is comprised of "simplex" autism families: those with at least two children, only one of whom has autism spectrum disorder or ASD. In such cases, disease causation has been previously linked to de novo, or spontaneously occurring, gene mutations.
The new research reached its conclusions based on full exome sequencing of each family member's genome. The exome is the portion of the genome – less than 2% of the total -- in which DNA encodes proteins. By studying simplex families, researchers seek to discover, among other things, the fraction of autism caused by gene mutations that are not inherited.
This study focused on small-scale de novo mutations – changes in DNA as small as a single DNA "letter" relative to the normal sequence, and small insertions or deletions as large as 10 or 15 letters. In the majority of children with small de novo mutations those mutations traced to the father's germ cells (sperm), in an age-dependent manner. Thus, the children of older parents were more likely to have the sorts of mutations that can cause autism.
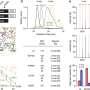
There are many types of gene mutations, some more likely to result in biological dysfunction than others. One of the most important findings of this study concerned the type of small-scale de novo mutations most likely to play a causative role in autism. "We found that those small-scale de novo changes that disrupt genes – in other words, those that disrupt the production of a normal length protein -- are twice as frequent in affected vs. unaffected children," says CSHL Assistant Professor Ivan Iossifov, a quantitative biologist who is the study's other co-first author. (In technical terms, these "disrupting" mutations include frame-shift, splice-site, and nonsense mutations, but not missense mutations.)
The estimated incidence of ASD is about 1 in 100 live births. De novo mutations are fairly common, 100 occurring on average in every child born. But only one such mutation per child typically occurs in a chromosomal region encoding a gene, and the odds that the mutation actually disrupts a gene is lower still. Finally, the odds that the gene disrupted is a gene important in the development of the brain lowers the risk of an autism-causing mutation further.
Wigler's team estimates there are 350 to 400 such vulnerable genes out of the full human complement of about 20,000 genes. "It is this 2:1 differential between the incidence of gene-disrupting mutations seen in affected vs. unaffected siblings, and the number of recurrent and total targets of gene disruption found in our study and recent similar studies, that is the basis of our estimate a total of between 350 and 400 autism susceptibility genes," Iossifov adds. Only about 10% of autism can be explained by mutations of the type the authors detected, which the authors reason is a large underestimate.
While the CSHL study identified about 60 previously unidentified autism candidate genes, and confirmed the role of a number of others previously identified, it was the linkage with the product of the Fragile-X gene that is perhaps its most intriguing result. A collaboration with Drs. Robert and Jennifer Darnell of Rockefeller University was critical in this regard. The Darnells had published extensively on FMRP, positing its biological mechanism of action in relation to autism.
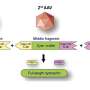
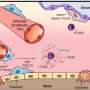
To clarify, the association discussed in the new paper is not with the FMR1 gene itself, but rather genes that are "downstream" of it. These are genes whose products FMR1 regulates – and regulates via its encoding of the protein FMRP. Once generated, FMRP binds to certain RNA messages (mRNAs) produced by other genes. When FMRP binds to these mRNAs, it prevents them from properly engaging the ribosome to generate proteins.
More information: Iossifov et al.: "De Novo Gene Disruptions in Children on the Autistic Spectrum" appears online ahead of print in Neuron on April 25, 2012.