Online platform simulates how the body defends itself
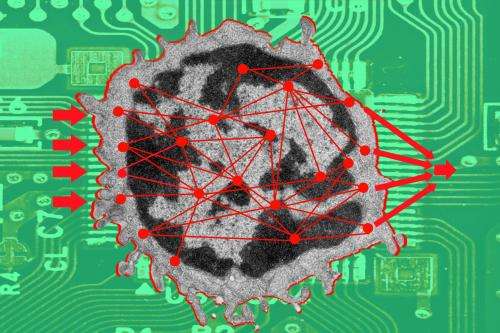
A platform that simulates how the body defends itself: The T cells of the immune system decide whether to trigger an immune response against foreign substances. Since December 2013 scientists from around the world can use the "virtual T cell" to test for themselves what happens in the blood cell when receptor proteins are activated on the surface.
Prof. Dr. Wolfgang Schamel from the Institute of Biology III, Facutly of Biology, the Cluster of Excellence BIOSS Centre for Biological Signalling Studies and the Center of Chromic Immunodeficiency of the University of Freiburg is coordinating the European Union-funded project SYBILLA, "Systems Biology of T-Cell Activation in Health and Disease." This consortium of 17 partners from science and industry has been working since 2008 to understand the T cell as a system. Now the findings of the project are available to the public on an interactive platform. Simulating the signaling pathways in the cell enables researchers to develop new therapeutic approaches for cancer, autoimmune diseases, and infectious diseases.
The T cell is activated by vaccines, allergens, bacteria, or viruses. The T cell receptor identifies these foreign substances and sets off intracellular signaling cascades. This response is then modified by many further receptors. In the end, the network of signaling proteins results in cell division, growth, or the release of messengers that guide other cells of the immune system. The network initiates the attack on the foreign substances. Sometimes, however, the process of activation goes awry: The T cells mistakenly attack the body's own cells, as in autoimmune diseases, or they ignore harmful cells like cancer cells.
The online platform developed by Dr. Utz-Uwe Haus and Prof. Dr. Robert Weismantel from the Department of Mathematics of ETH Zurich in collaboration with Dr. Jonathan Lindquist and Prof. Dr. Burkhart Schraven from the Institute of Molecular and Clinical Immunology of the University of Magdeburg and the Helmholtz Center for Infection Research in Braunschweig allows researchers to click through the signaling network of the T cells: Users can switch on twelve receptors, including the T cell receptor, identify the signals on the surface of other cells, or bind messengers.
The mathematical model then calculates the behavior of the network out of the 403 elements in the system. The result is a combination of the activity of 52 proteins that predict what will happen with the cell: They change the way in which the DNA is read and thus also that which the cell produces. Now researchers can find weak points for active substances that could be used to treat immune diseases or cancer by switching on and off particular signals in the model. Every protein and every interaction between proteins is described in detail in the network, backed up with references to publications. In addition, users can even extend the model themselves to include further signaling proteins.










.jpg)







