Researchers map genome-wide changes that drive T cell maturation and exhaustion
In a bid to better understand the gene expression patterns that control T cell activity, researchers at the La Jolla Institute for Allergy and Immunology mapped genome-wide changes in chromatin accessibility as T cells respond to acute and chronic virus infections. Their findings, published in the Dec. 20, 2016 issue of Immunity, shed light on the molecular mechanisms that determine the fate of T lymphocytes and open new approaches to clinical intervention strategies to modulate T cell activity and improve immune function.
"Identifying the different factors that determine different T cell states and therefore their function helps us understand if T cells will be able or not to fight viral infections or tumor growth, and if they will be able or not to provide long-term protection," says the study's first author James Scott-Browne, a postdoctoral fellow in the laboratory of Anjana Rao, a professor in the Division of Signaling and Gene Expression. "We may be able to revert the exhaustion phenotype of T cells and render them better able to fight tumors or chronic viral infections such as HIV, or generate better memory cells in response to vaccines."
When viruses invade or cells turn malignant, the immune system mobilizes a small cohort of naïve or immature CD8 T cells, a crucial subdivision of the immune system charged with killing virus-infected and cancerous cells. Upon activation, they mature and proliferate exponentially into highly specific effector T cells that eliminate virus-infected or otherwise compromised cells. After their job is done, most effector T cells die leaving behind only a small contingent of memory T cell that confer long-term protection.
In the face of chronic viral infections such as hepatitis and HIV as well as certain types of cancers, however, activated CD8 T cells are unable to gain the upper hand and clear the threat. As a result, CD8 T cells start to express inhibitory cell surface receptors that transmit inhibitory signals into the cell establishing a negative feedback loop. The mechanism is designed to prevent excessive immune responses from taking hold but it leaves CD8 T cells unable to fight foreign invaders effectively and forces them into a state known as "T cell exhaustion."
In earlier work, Rao and her team had pinpointed a transcription factor known as NFAT as the molecular linchpin that orchestrates T cell activation and exhaustion. When the T cell receptor on the surface of CD8 T cells recognizes a foreign protein, it kicks off a signaling cascade that culminates in the activation of NFAT and its partner AP-1.
Together, the pair binds to regulatory regions in the genome and initiates a genetic program that activates T cells and readies them to fight cancer and viral infections. When acting on its own, NFAT shifts the equilibrium from an activated to an exhausted state by binding to a different subset of regulatory regions within the genome, impairing the immune system's response to tumors and infection.
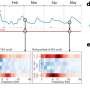
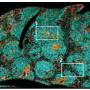
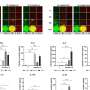
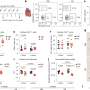
The current study expands the previous experiments, which were largely based on lab-grown T cells, to T cells isolated from mice with acute or chronic viral infections. It centered on a powerful methodology known as ATAC-seq, which pinpoints "open" or accessible stretches of chromatin. Chromatin is the sum total of genomic DNA and all associated proteins, which not only packages and condenses DNA but also helps control gene expression by giving or denying access to transcription factors. Knowing which regulatory sites in the genome are open for business allows scientists to conclude which transcription factors play a role in certain biological processes.
"We showed that when naïve cells are transformed into effector cells, there are big changes in the regions of chromatin near the genes that determine an "activated fate", explains co-lead author Renata Pereira, formerly a postdoctoral researcher in the Rao laboratory, and now an assistant professor at the Universidade Federal do Rio de Janeiro in Brazil. "In contrast the chromatin structure of effector cells is quite similar to that in memory or exhausted cells, suggesting that the differences in the functions of these cell types depend mostly on the action of transcription factors that bind the already open chromatin regions. So transcription factors could be a more interesting target to modulate the function of T cells than proteins that modulate if the chromatin is more or less accessible."
The work was funded by the National Institutes of Health (R01 AI40127) (to A.R.). the Damon Runyon Cancer Research Foundation, and the Pew Latin American Fellows Program in the Biomedical Sciences.