Cells study helping to crack the code to Alzheimer's disease
A study led by researchers at Monash University has opened up new hope for diagnosing and treating Alzheimer's disease.
Alzheimer's disease is the most common form of dementia in older people and, as there are no effective treatments, is one of the leading contributors to the global disease burden.
Various genes have been implicated in the changes that happen in the brains of Alzheimer's patients. What is not known is how the activity of the genes—called gene expression—affects the many different cells of the brain.
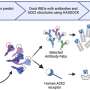
The study, led by Monash University's Professor Jose Polo, from the Monash Biomedicine Discovery Institute and the Australian Regenerative Medicine Institute, focused on how individual cell types in the brain contribute to Alzheimer's disease.
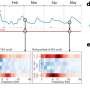
Through a series of complex tests, they looked at patterns in gene expression in specific cells and how changes in specific cell subpopulations are associated with Alzheimer's.
The researchers were excited to make a number of key discoveries into understanding Alzheimer's disease that require further study.
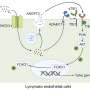
This included the role of disease gene networks, genes working together, in the human brain, pathways to genetic susceptibility, subcellular changes in areas of the brain, and the role of myelination in Alzheimer's disease pathogenesis.
The study was co-led with researchers from Duke-National University of Singapore and in collaboration with the University of Melbourne, Florey Institute, the University of Western Australia and the Harry Perkins Institute of Medical Research. This is an excellent outcome from the Monash International Network of Excellence program, that is designed to bring together new international teams working towards the solution of big problems.
In a paper published in Nature Neuroscience on Tuesday, November 26, the researchers said: "although our study contributes significantly to our understanding of the transcriptional changes underpinning changes in brain cells in Alzheimer's disease, examining larger patient cohorts in the future will enable assessment of the effects and relative contribution of underlying genetic factors to the described cellular and transcriptomic changes in disease.
"We anticipate that our resource will stimulate and allow further discoveries in many different areas, as exemplified by our work."
Professor Polo said the findings are an exciting step in eventually diagnosing and treating Alzheimer's disease.
"Alzheimer's disease is an increasingly prevalent cause of dementia in ageing societies like Australia and Singapore. Despite billions of funds poured in, an effective drug has yet to be discovered," the lead researchers said.
"In order to make advancements, there is a need to focus on other cell types in the brain aside from neurons—the main cell type in the brain.
A key researcher in the study, Dr. Alexandra Grubman, from Monash University's Australian Regenerative Medicine Institute and a NHMRC-ARC Dementia Fellow, is excited at the new therapeutic possibilities the study has opened.
"Perhaps the answer to treating Alzheimer's lies in understanding how these non-neuronal cells are affected during disease," she said.
This multinational team will now conduct further research on genes that can potentially be treated with medical drugs.
More information: A single-cell atlas of entorhinal cortex from individuals with Alzheimer's disease reveals cell-type-specific gene expression regulation, Nature Neuroscience (2019). DOI: 10.1038/s41593-019-0539-4 , nature.com/articles/s41593-019-0539-4