New mechanism by which senescent cells turn on genes encoding for tumor-regulating factors

Scientists at The Wistar Institute identified a new mechanism of transcriptional control of cellular senescence that drives the release of inflammatory molecules that influence tumor development through altering the surrounding microenvironment. The study, published in Nature Cell Biology, reports that methyltransferase-like 3 (METTL3) and 14 (METTL14) proteins moonlight as transcriptional regulators that allow for establishment of the senescence-associated secretory phenotype (SASP).
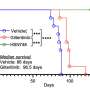
Cellular senescence is a stable state of growth arrest in which cells stop dividing but remain viable and produce an array of inflammatory and growth-promoting molecules collectively defined as SASP. These molecules account for the complex crosstalk between senescent cells and neighboring cells and the effect of cellular senescence in various physiological processes and diseases. Although senescence is regarded as a potent barrier for tumor development, the SASP plays a stage-dependent role during tumor development, mediating the clearance of premalignant lesions during initiation and promoting the growth of established tumors.
"Senescent cells undergo widespread changes in gene expression needed to adapt their phenotype and functions," said Rugang Zhang, Ph.D., deputy director of The Wistar Institute Cancer Center, Christopher M. Davis Professor and leader of the Immunology, Microenvironment & Metastasis Program. "We pointed out a new mechanism that allows cells to turn on a set of genes encoding for the SASP molecules and may potentially be targeted to inhibit this aspect of senescence while preserving its antitumor function."
Zhang, who is senior author on the study, and his team focused on METTL3 and METTL14, proteins known for chemically modifying messenger RNA to regulate its function. They found a new role of these proteins in senescence and regulation of gene expression that is independent of their RNA-modifying function.
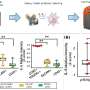
Depleting cells of METTL3 and METTL14, researchers observed reduced expression of SASP genes, such as inflammatory cytokines, but no effect on cell cycle arrest or other markers of senescence, indicating that decrease in SASP is not an indirect consequence of overall senescence inhibition.
"Our results indicate that METTL3 and METTL14 promote expression of SASP genes, in accordance with other studies that revealed an oncogenic role for these two proteins," said Pingyu Liu, Ph.D., first author of the study and a staff scientist in the Zhang Lab.
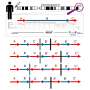
The team further analyzed the association of METTL3 and METTL14 with DNA, comparing senescent and control cells. While the two proteins are found together on DNA in control cells, in senescent cells they have different distribution patterns, whereby METTL3 tends to sit upstream of SASP genes, near the transcription start site, while METTL14 binds away from gene bodies, on regulatory elements called enhancers.
Researchers demonstrated that through this positioning pattern and interacting with each other, METTL3 and METTL14 bring closer together two DNA sequences that in non-senescent cells are distant, allowing the formation of promoter-enhancer chromatin loops. As a consequence, expression of the SASP genes is turned on.
"Although we focused on senescence, we envision that the transcription-regulating function of METTL3 and METTL14 may be involved in many other biological processes beyond our current study," concluded Zhang.
More information: m6A-independent genome-wide METTL3 and METTL14 redistribution drives senescence-associated secretory phenotype, Nature Cell Biology (2021). DOI: 10.1038/s41556-021-00656-3