Protein may protect against neurodegenerative diseases
Cells translate their genetic material at rapid rates with exquisite precision to reproduce, repair damage or even combat disease. But the process can deregulate and give rise to disease. Byproducts of errant processes can build up like gunk in the gears, especially around neurons, breaking down the repair mechanisms and causing further damage and even neurodegenerative disorders such as Alzheimer's disease.
Now, an international research team may have found that a protein implicated in tumor growth may be able to help regulate awry cellular translation and protect against neuronal decay. They published their results on July 13 in the Cell Reports.
"Researchers have begun to understand that age-related neurodegenerative diseases may be caused by slow but steady accumulation of toxic peptide products, which leads to death of neurons, such as beta-amyloid plaques causing Alzheimer's disease," said lead paper author Katsura Asano, professor in Hiroshima University's Graduate School of Integrated Sciences for Life and also in Division of Biology, Kansas State University. "Repeat-associated non-AUG (RAN) translation is one of the mechanisms that generates such toxic products."
When replicating their genetic material, cells look for specifically ordered markers that signal the spot where they should start and end the copy to make a specific protein. The signal is typically "AUG," but RAN translation doesn't need this signal and can begin processing at other points. The problem is that RAN translation can end up copying bits of repeated genetic information that become the toxic buildup that leads to neurodegeneration.
"Taking advantage of our knowledge on translation, we examined whether the translational regulatory protein 5MP can suppress RAN translation, which would also suppress the production of the toxic repeat peptide products," Asano said.
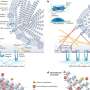
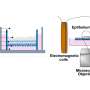
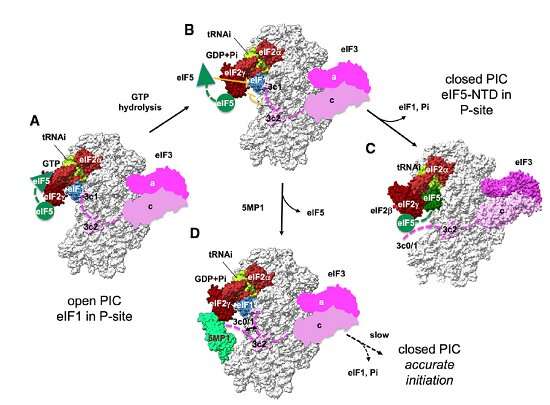
In diseased conditions, 5MP is a protein that can potentially transform a healthy cell into a tumor cell. In healthy conditions, 5MP mimics a protein involved in regulating RAN translation. The researchers used electron microscopy and computer-directed modeling to reveal the structure of the preinitiation complex of molecules that assemble prior to RAN translation beginning. They found that 5MP competes with the protein it mimics in human cells and, when it wins, it reduces RAN translation and its toxic byproducts.
To better understand how this finding translates to disease outcomes, the researchers engineered flies with fragile X-associated tremor ataxia syndrome, a neurodegenerative disorder. They found that increasing the levels of 5MP in the affected flies repressed neuronal toxicity and enhanced their lifespan.
"Taken together, these data suggest that modulation of 5MP levels could be a viable therapeutic target by which to selectively reduce RAN translation in repeat expansion disorders," Asano said. "More studies on 5MP and the mechanism of translation can greatly contribute to the understanding and care of neurodegenerative diseases."
More information: Chingakham Ranjit Singh et al, Human oncoprotein 5MP suppresses general and repeat-associated non-AUG translation via eIF3 by a common mechanism, Cell Reports (2021). DOI: 10.1016/j.celrep.2021.109376