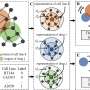
New algorithm dramatically increases the speed of identifying two cancer drugs that work synergistically

An algorithm that can speed up by years the ability to identify from among thousands of possibilities, two or more drugs that work synergistically against a problem like cancer or a viral infection has been developed by bioinformatics experts.
The new algorithm enables investigators to use large existing databases with information about how one cancer drug changed the gene expression of a particular breast cancer cell line, and how well it killed the cell, then mathematically combine those results with the impact of another drug to see if they could work better together, says Dr. Richard McIndoe, director of the Center for Biotechnology and Genomic Medicine at the Medical College of Georgia.
While the algorithm does not immediately make available the kind of information that would set a clinical trial in motion, it does speed up the path to the trials, he says.
"The idea is we ultimately want to find these synergistic drug combinations that will hopefully help patients with cancer," McIndoe says. "For researchers it becomes a particularly faster way to find those synergistic combinations, without having to screen one drug at a time, which is really not feasible."
Drug combination therapies can improve drug efficiency, reduce drug dosage (and related toxicity) and overcome drug resistance in cancer treatments," the investigators write in the journal PLOS ONE, and is becoming an important tool in cancer treatment.
"It's not uncommon for the cancer to become resistant to chemotherapy drugs so one of the ways that clinicians try to get around that is using combinations, two chemotherapy drugs together," McIndoe says. "The likelihood that you will develop resistance to both of them simultaneously is lower than if you had just one."
But given the number of drugs and drug combinations available, there are not efficient, effective ways to identify the best combinations, the investigators say.
And, not all combinations are beneficial, in fact one drug can actually work as an antagonist against the other, effectively blocking or at least reducing its therapeutic impact. The right combination will, by contrast, enhance the impact of the treatment, McIndoe says, which means together they are better at killing cancer cells-.
The algorithm also enables synergy between scientists by enabling easily sharing findings which enables even more drugs and cell lines to be evaluated and the database of effective combinations against specific cancers to grow more rapidly.
"The tricky part is how do you determine which drug combinations have a synergistic effect," he says.
Existing approaches to finding the right combination include huge, automated stations where different drug combinations are put with a specific cancer cell line to see what happens. But the list of drugs is long and potential combinations even longer, McIndoe says.
Another approach is to put drugs together based on what is known about their respective modes of action, which still amounts to a lot of drugs and drug combinations, another slow and costly process, he says.
There are huge existing databases of cell lines which have been treated with one drug to look at the impact on gene expression, before and after treatment, including the Library of Integrated Network-based Cellular Signatures project, to help streamline the large-scale studies like the MCG investigators wanted to do.
They focused on 57 randomly selected chemotherapy drugs used in the database, looking in detail at the molecular changes each drug produced and tying that to growth rate, meaning how much cancer cell killing the drug produced, then devised a mathematical representation of the molecular changes and the amount of killing for each.
"Since we have all the single drug effects, we can mathematically combine two drugs based on their molecular changes," McIndoe says.
All told there were 1,596 combinations of the 57 cancer drugs they studied. Their algorithm picked 30 top drug combinations and eight were confirmed using a standard statistical model called ZIP, a result far better than chance and far less costly and time consuming than testing the large number of potential drug combinations, McIndoe and his colleagues write.
Follow up tests in the lab, where they treated the cancer cell lines with the proposed synergistic combinations, further confirmed their synergy.
He notes that to screen all 1,596 combinations would take about three years using standard approaches, compared to about eight weeks with their algorithm.
"What we were asking for this paper is can we use that gene expression data to come up with a way to prioritize which drugs would have the highest probability of being synergistic when you put them together," McIndoe says.
Gene expression changes or mutations from things like environmental exposures or even random mistakes, are a major cause in cancer's development and spread. Different classes of cancer drugs work in different ways to damage the DNA of cancer cells and prevent or at least slow their reproduction.
"When you hit a cell with anything, such as a drug or a nutrient change, the cell responds," he says. "It responds in a way where it is going to start to change its gene expression profile."
The adjustments likely result from the cancer cell trying to survive, like working to turn on some alternative signaling pathway to enable growth because the drug has shut the usual one down. But if the drug works, the cell will die, so McIndoe and his colleagues used kill rate as their important bottom line.
The investigators note that only a small number of the 57 drugs appeared to have no effect on the cancer cell line, and those drugs were not among those found to have a synergistic impact, but synergy is still possible when they are combined with the right partner, McIndoe says.
The algorithm could similarly be used to more readily identify the best drug combinations for other problems like bacterial, fungal and viral infections, they write.
Next steps include looking at other breast cancer cell lines treated with those same 57 drugs, looking again at their molecular changes in response and whether those changes are the same in different cell lines. McIndoe also wants to create a database where other investigators can easily upload the impact on gene expression and growth rate for their studies and take the important next step in lab animal studies to see if the synergy holds in an intact tumor, for example.
The first author on the study is Dr. Jiaqi Li, a former graduate student with McIndoe who took on the project for his thesis to use bioinformatics tools to push science forward. Li is now a research associate at the MCG Center for Biotechnology and Genomic Medicine. Coauthor Dr. Hongyan Xu is a human population geneticist and biostatistician in the MCG Department of Population Health Sciences.
More information: A novel network based linear model for prioritization of synergistic drug combinations, PLoS ONE (2022). journals.plos.org/plosone/arti … journal.pone.0266382