Interdisciplinary research team uses AI to quickly discover personalized cancer cures

LSU researchers have built a cancer drug discovery engine powered by AI that soon could match any type of cancer, based on a small cell sample from a patient, with the drug most likely to cure that cancer.
Surgical oncologist Dr. John Stewart, director of the LSU Health New Orleans/LCMC Health Cancer Center said, "LSU's work to leverage AI for precision medicine will allow us to devise new molecular-driven treatments for every patient."
An interdisciplinary LSU research team with members in the School of Veterinary Medicine, College of Science, College of Engineering and the Center for Computation & Technology is using artificial intelligence, or AI, to discover personalized cures for cancer more quickly and affordably.
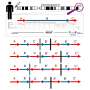
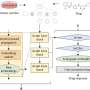
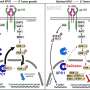
Developing algorithms originally designed to map complex social networks, such as those used by Facebook, the researchers have created a cancer drug discovery engine called CancerOmicsNet. It combines three-dimensional graph representations of vast molecular datasets, including cancer cell lines, drug compounds, as well as protein-protein interactions. Once analyzed and interconnected by AI, they form a much clearer picture of how a specific cancer would respond to a specific drug, removing much of the guesswork in current oncology.
"Cancer is a very complex condition," said Michal Brylinski, associate professor of computational biology at LSU.
"There are hundreds of different types of cancer, and every type of tissue has some sort of cancer—and there are many different subtypes—so there's huge diversity in possible cancer targets. Meanwhile, pharmaceutical companies have developed hundreds of cancer drugs, but not all drugs are potent against all cancers. Today, you cannot say 'This drug is going to work.' But with CancerOmicsNet, we can make much more accurate predictions, and fast."
The researchers have completed an initial study of breast, prostate and pancreatic cell lines, the human cancer cells available for medical research, which are among the most aggressive and difficult to treat. They trained their AI to recognize meaningful connections between specific cancers and cancer drugs that control kinase production in the body.
Kinase is an enzyme, a biological catalyst for cell communication and cell growth. With drugs that lower kinase activity, cancer cell growth can be suppressed. But how effective a kinase-inhibiting drug will be for a certain type of cancer involves considerable guesswork. Also, the most effective drug available on the market is sometimes not considered because it's only known to work for other diseases.
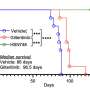
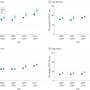
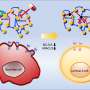
CancerOmicsNet can overcome these challenges. Like molecular speed dating, the researchers' AI can quickly match cancer cell lines with the drugs likely to be their most "toxic partners" or the most opposite in the way that matters most, namely their gene expression profile. After testing six such combinations, all previously unknown before being identified by the team's AI, the researchers found something extraordinary—four of the six combinations worked. Cancer growth was slowed, even reversed.
"Normally, if you select six combinations at random, nothing is going to work," Brylinski said. "I was hoping, praying at least one would work, but instead we had four."
"These predictions would have been virtually impossible for people to make, but complex problems are naturally a very good target for AI," Brylinski continued.
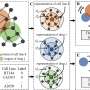
"Our biggest challenge was to figure out how to visually represent a cancer as a graph or network, where each node in the graph is a protein. We have about 19,000 of these proteins in human cells, including cancer cells, and those proteins interact with each other through edges. You can imagine how complicated these relationships are compared to simple gene signatures, which is how people traditionally have approached the problem of discovering effective cancer drugs."
Deep learning, considered the most human-like type of AI since it's capable of independent "thought" and decision-making, has already started to significantly impact biological research and biomedical applications. This is due to AI's ability to extract patterns from highly dimensional and complex data, such as similarities and differences between drug molecules and cancer cells.
"In the state of Louisiana, we have some of the poorest outcomes of cancer in the entire country, and a higher incidence of cancer in younger patients," Dr. Stewart said. "This is why the advent of precision medicine is so important—because it facilitates targeted therapies for cancer."
Early in Dr. Stewart's career, oncologists thought of cancer as specific to its location, such as lung cancer in one patient being similar to lung cancer in another patient.
"We assumed molecular consistency," Dr. Stewart said. "But what we understand now is that cancer is several hundred diseases with differences across the genetic landscape. Genomic testing in combination with AI might reveal that a colon tumor has a similar genetic mutation profile to a breast cancer, for example, and is susceptible to the same targeted therapy."
This could lead to more effective cancer treatments for everyone in Louisiana—regardless of individual differences—and help overcome health disparities, Dr. Stewart argues.
"LSU's work to leverage AI for precision medicine will not only allow us to devise new molecular-driven treatments for every patient, but also allow us to understand how to provide preventative services to patients in rural communities," he said.
The ongoing LSU collaboration involves both dry and wet lab research conducted on the main Baton Rouge campus. The AI development and computational biology, or "dry" theory, is followed by hands-on studies of actual cancer cells in the LSU School of Veterinary Medicine.
The wet lab work is led by Assistant Professor of Research Brent Stanfield who has significant experience with using viruses to treat various diseases, which is perhaps an odd but interesting approach, and to manipulate cells—leveraging the power of viruses to enter cells quickly and efficiently, but with a harmless or beneficial end result.
"In this study, we used a lentivirus to deliver a fluorescent reporter to each cancer cell," Stanfield said. "This didn't physiologically change the cells, it just acted as a courier to label the cells to make them easier to see with a microscope, so it became easier for us to count the cancer cells before and after we applied each drug."
"I love this work," Stanfield continued. "I wake up every day to do this. I'm really grateful LSU has the cutting-edge research infrastructure it has with these big center grants because it's really transforming what kind of research we can do right here on campus with high-throughput and high-capacity screening techniques."
Next, the researchers are going to expand the scope of their study beyond kinase-inhibiting drugs, and also use their AI to discover effective combinations of cancer drugs at scale.
"We can train our AI to select two drugs to treat the same cancer, so the drugs also have a synergistic effect and amplify each other," Brylinski said. "This can be super-beneficial to patients because, if we can give drugs in lower doses, patients suffer less side effects."
Three former LSU students are part of the interdisciplinary CancerOmicsNet team: 2019 electrical engineering graduate Limeng Pu, who now serves as IT consultant at the LSU Center for Computation & Technology and helped develop the cancer drug discovery engine's graph-based convolutional neural network, or how the AI "thinks"; Manali Singha, who just graduated with a Ph.D. in biological sciences and helped conceptualize biological data utilization, data acquisition and data integration, or preparing input for the AI; and Ifeanyi "Kingsley" Uche, who assisted with the wet lab work and recently graduated from the LSU School of Veterinary Medicine to pursue medical school at LSU Health New Orleans.
"While people have been generating and collecting all sorts of biological and chemical data for decades, it's only with the advance of high-performance computing—both hardware and software—and deep learning and AI that we now have the capacity to deal with this data in effective and useful ways to improve human health," Pu said.
"Remember, the cost of developing drugs is very high. The research and development in the earliest stages, long before clinical trials, can cost millions of dollars. By using AI, we can effectively reduce the cost of drug development by 40 percent. We can bring down the price and the time it takes."
This reasoning is familiar to Brylinski, who also is a member of the LSU DeepDrug team, which developed an AI drug discovery platform that was licensed to Skymount Medical in 2020 and used to rapidly find new treatments for COVID-19—technology that recently garnered the American Business Association's Silver Stevie award.
"This technology is very similar, while our focus here is on tumor profiling," Brylinski said. "In the future, doctors could potentially take a cancer sample from a patient and run simple, low-cost genomic testing—no more than a few hundred dollars—and use our AI technology to select the most effective drug or combination of drugs."