This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Study on DNA transcription uncovers links to neurodegenerative disease
In a first-of-its-kind study, Arizona State University Professor Michael Lynch joins a multi-institute group of researchers to investigate transcription error rates in human cells and the underlying mechanisms affecting them.
Transcription is the process of copying DNA to RNA. The accuracy of transcription processes varies widely among species, across cell types and within distinct regions of the genome, with profound consequences for health and disease. DNA transcription is a crucial process in the expression of genetic information, as it converts the information stored in DNA into messenger RNA, which can then be translated into proteins.
The research sheds new light on a foundational process in biology. The results also suggest that high rates of transcription error, observed in specific classes of neurons, are a potential source of neurodegenerative diseases, including Alzheimer's disease.
"Although we have recently made substantial progress on estimating error rates at the transcriptional level, the next challenge is to establish the connection of such errors with cell health," Lynch says.
The research results appear in the current issue of the journal Proceedings of the National Academy of Sciences.
Professor Lynch directs the Biodesign Center for Mechanisms of Evolution and is a professor in the School of Life Sciences at ASU. He is one of the world's leading quantitative geneticists, whose research focuses on uncovering the mechanisms driving evolution at the genomic, cellular and organismic levels. He has recently been honored with an ASU Regents Professorship.
The new study analyzes transcription errors in human embryonic stem cells and in mice to unveil the molecular mechanisms governing transcriptional accuracy. The research provides the first estimate of transcriptional error rates in human cells and identifies various genetic and epigenetic factors responsible.
Reading DNA is fundamental
The foundations of life hinge on the precise replication and transcription of DNA and the translation of the resultant messenger RNAs. These processes are responsible for accurately passing down and expressing our genetic information, and their fidelity is crucial for maintaining the stability of our genetic code. Despite their importance, the molecular mechanisms behind the faithful transcription of DNA remain largely unknown.
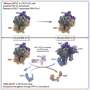
During transcription, the genetic information stored in a gene's DNA sequence is copied into a molecule of messenger RNA (mRNA), which then carries the information out of the nucleus and into the cytoplasm where it can be translated into a functional protein. Transcription errors occur during the process of copying genetic information from DNA to RNA, one of the key steps in gene expression.
These errors can arise from DNA damage, incorrect recognition of the DNA template by the gene-reading mechanism (known as RNA polymerase) or problems with the repair mechanisms that correct errors in the transcription process.
Inaccurate transcription can produce truncated or altered proteins that are unable to perform their normal functions, leading to disease.
Uncovering error
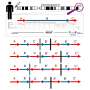
Several factors exert a profound influence on rates of transcription error. Some genes are more faithfully transcribed than others, which can be a consequence of their relative length or complexity. Genes are sequences composed of DNA's 4 nucleotides, labeled A, T, C and G.
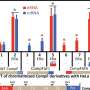
The study demonstrates they are not transcribed with equal reliability, as A and G transcriptions tend to be more error prone.
The study also reveals that different types of RNA polymerase, the machinery responsible for proofreading DNA during transcription, have significantly differing rates of reliability. The error rate not only differs between types of polymerase, but also between classes of genes being transcribed and even between specific regions of these genes.
Another key factor of accuracy is the rate of transcription. Just as a proofreader is more likely to make mistakes if they race through a page of text, ultra-rapid DNA reading by fast RNA polymerases are more likely to produce errors in transcription.
There are also differences in the behavior and effectiveness of DNA repair proteins, which can fix mistakes in transcription after they have occurred. A new role for one such protein, known as BRCA1, is reported in the study. In addition to BRCA1's role in repairing DNA damage and preventing it from accumulating across the genome, the study indicates this invaluable protein appears to improve transcription fidelity.
Mutations in the BRCA1 gene, which codes for this error-correcting protein, have long been associated with a range of serious health issues, particularly breast cancer and ovarian cancer. BRCA1 mutations have also been linked to other health conditions, including pancreatic cancer, melanoma and fallopian tube cancer.
Vulnerabilities in the brain
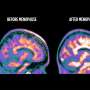
A mouse model was developed to probe which cell types are most susceptible to producing misfolded proteins due to transcription errors. Neuronal cell types associated with Alzheimer's disease display comparatively high transcriptional error rates. One of the effects of this appears to be the generation of a toxic protein form called APP, a precursor to the amyloid plaques that accumulate and cloud the intercellular spaces of the brain and which are a hallmark of Alzheimer's disease.
Cells and tissues most prone to transcription errors are identified in the study, revealing that neurons in two critical regions of the brain, CA1 and dentate gyrus, are particularly disposed to DNA alterations or transcriptional mutagenesis. The finding supports the hypothesis that transcription errors contribute to Alzheimer's disease and other potentially devastating effects in the brain.
Such protein aberrations produced by transcription errors may be culprits in other neurodegenerative diseases, including Parkinson's disease, amyotrophic lateral sclerosis and frontotemporal dementia.
The foundation of life lies in the precise replication, transcription and translation of DNA but knowledge about the mechanisms that control the accuracy of transcription remains limited. Ongoing research of this kind will deepen understanding of processes at the heart of biology and may advance new approaches to currently intractable afflictions, such as Alzheimer's disease.
More information: Claire Chung et al, The fidelity of transcription in human cells, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2210038120