Adult leukaemia can be caused by gene implicated in breast cancer and obesity
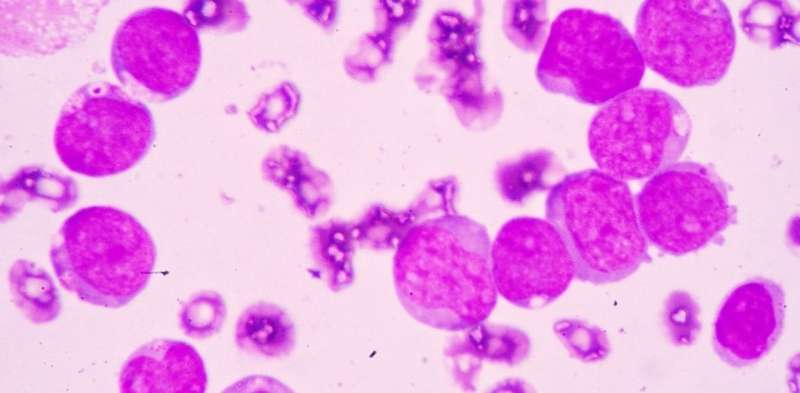
When people think of leukaemia, they usually think of blood cancers that affect children. These mostly come under the category of acute lymphoblastic leukaemia – or ALL – and are different to the group of blood cancers which predominantly affect adults over the age of 60, known as acute myeloid leukaemia (AML).
AML accounts for about 90% of all leukaemias in adults, though it affects some children too. With some 3,000 new cases each year in the UK alone, it is tougher to treat than ALL.
Where advances in ALL treatment have raised survival rates to around 90% over the past several decades, the rates for surviving the less well researched AML are more like 65%. Older adults respond least well to treatment, with only 5% of over-65s surviving more than five years.
I am therefore pleased to report a promising discovery. Work in which I have been involved has shown that a particular gene can play a critical role in the development of the disease. This could be the precursor to a breakthrough that could be life-saving for patients.
Cells and treatments
Your bone marrow contains stem cells which divide and differentiate into red blood cells and the main groups of white blood cells – myeloid cells, neutrophils and lymphocytes. Normally this happens in a very controlled manner, ensuring you have all the red blood cells needed to carry oxygen around your body, and all the white blood cells needed to fight off infections.
In AML too many immature myeloid cells are produced too quickly by the bone marrow. They are mutant cells which don't mature, meaning they fail to defend against infection.
For this reason, early signs of AML include flu-like symptoms, aches and pains in the joints, and rapid weight loss. As the abnormal cells build up inside the bone marrow or the blood they grow and divide aggressively. Left untreated, AML patients can have only weeks to live.
Patients are normally treated by two stages of chemotherapy over a few months: an induction phase which reduces the number of cancer cells to undetectable levels, then a consolidation phase to kill any cancerous cells hiding in the body. Patients often also receive a bone marrow transplant, effectively giving them a new immune system.
Treating AML is complicated by patients generally being older, since they tolerate the intensive chemotherapy less well. In many cases, they receive some treatment and end up only living a few months. Better understanding the mutations to develop more targeted and less harsh treatments looks like the key to improving survival.
Step forward, PTPN1
A number of mutations are associated with AML, and often occur in combinations. It's these mixtures of mutations that are thought to cause the complex subtypes of cancers within the AML group. One common mutation is called Del20q. It involves the deletion of part of chromosome 20, one of the 23 pairs of chromosomes most humans have in all their cells.
It has long been suspected that genes on this part of the chromosome may function, either individually or together, to suppress cancer. Until recently, however, researchers have found it hard to say which genes are responsible.
One candidate is known as PTPN1, or protein tyrosine phosphatase, non-receptor type 1. First discovered in the late 1980s and linked to metabolic function, it is more famously known for its roles in breast cancer and type 2 diabetes. Its location on chromosome 20 has long made specialists suspect it could also be involved in AML.
It was shown recently that when you switch off the equivalent gene in mice, it leads to what are known as myeloproliferative neoplasm, which is the wider family of blood cancers of which AML is a member. In our new study, we have taken this a step forward: we have shown that if you delete this gene in older mice, it specifically gives rise to AML – and in a similar way to how the disease develops in older humans.
The previous study showed that PTPN1 is deleted from chromosome 20 in the cells of patients in around 17% of AML cases, which raises questions about the remaining majority of cases. We were able to show that deleting the mouse equivalent of PTPN1 activates a molecule called STAT3, which is important to regulating cell growth and division.
If a patient has too much STAT3, it leads to the generation of too many immature myeloid cells – that hallmark of AML I mentioned earlier. This is potentially a very useful finding for further studies into the genetics behind the disease: in the two other most common mutations linked to AML, which relate to a protein called JAK2 and a receptor called FLT-3, STAT3 is also over-activated. In all, STAT3 is relevant to maybe three quarters of all AML cases. Uncovering exactly how they relate looks critical to developing an eventual cure.
The future
In short, we're closing in on understanding the links between PTPN1, STAT3 and AML. A few years from now, as the cost of genome sequencing falls, it will become a question of identifying which combination of mutations has affected a patient and prescribing a treatment accordingly.
This treatment will probably be more bespoke chemotherapy for patients that can tolerate it, and perhaps gene editing using tools such as CRISPR for those that cannot. Doctors would edit the correct versions of genes like PTPN1 back into the patient's bone marrow, potentially restoring normal function and negating the often difficult search for a compatible bone marrow donor.
There is much research still to be done. We need to understand what PTPN1 is doing in healthy myeloid cells to grasp which processes are disturbed when it becomes deleted. The other big question is whether instead of getting deleted, PTPN1 sometimes more subtly mutates and how this relates to AML. Besides this, there are many other genes on the Del20q deletion that we need to better understand, too.
In the meantime, showing that removing PTPN1 leads to AML is an important piece of the puzzle. It brings the day closer when survival rates for AML make the same climb that we have seen in other kinds of leukaemia, and hopefully even beyond.
This article was originally published on The Conversation. Read the original article.![]()