Researchers discover insights into amyloids associated with Alzheimer's and type 2 diabetes
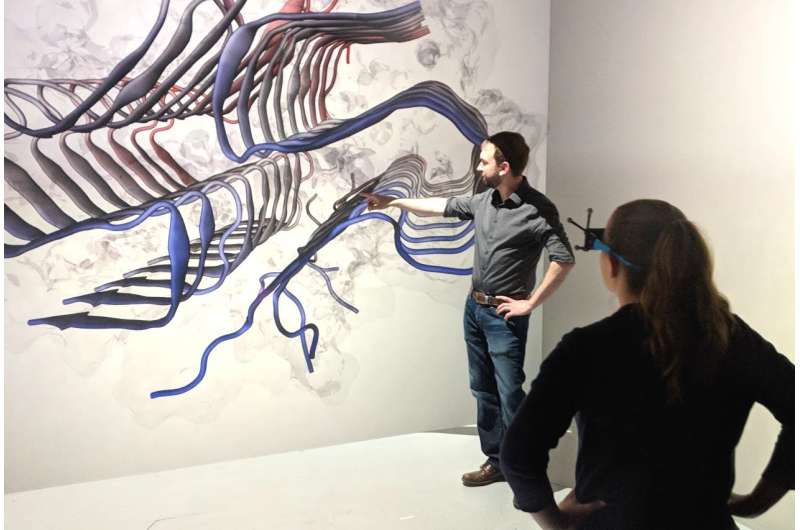

A Virginia Tech research team has discovered insights into the stabilizing forces of amyloid fibrils that are associated with Alzheimer's disease and type 2 diabetes.
These findings were recently published in the Journal of Molecular Biology.
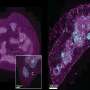
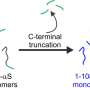
Amyloids are aggregates of proteins that form a shape that allows many copies of that protein to stick together to form fibrils. The accumulation of amyloid fibrils in the brain contributes to Alzheimer's disease, and the accumulation of amyloid fibrils in the pancreas contributes to type 2 diabetes by damaging cells that produce insulin.
Justin Lemkul, an assistant professor of biochemistry in the College of Agriculture and Life Sciences, and his team's research focuses on applying computer simulations to understand mechanisms of protein aggregation that are difficult or even impossible to recreate in a laboratory setting.
Elucidating the structure and stability of these amyloid fibrils is important for developing future anti-amyloid drug therapies.
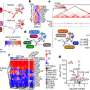
For this research, Lemkul's team performed the first-ever simulations of amyloid fibrils using a physical model that included electronic polarization to understand the forces stabilizing three amyloid-forming proteins observed in Alzheimer patients: microtubule-associated protein tau, amyloid β-peptide, and islet amyloid polypeptide (IAPP). IAPP is also associated with amyloid fibrils in type 2 diabetes patients.
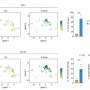
"We found that several amino acids in these three amyloid-forming proteins are particularly sensitive to small changes in their environment, particularly glycine, which plays a major role in stabilizing amyloid aggregates," said Lemkul, an affiliate of the Fralin Life Science Institute and Virginia Tech Center for Drug Discovery.
Anne Brown, an assistant professor in research and informatics, University Libraries, is a contributing author and performed the IAPP simulations and analysis for the paper.
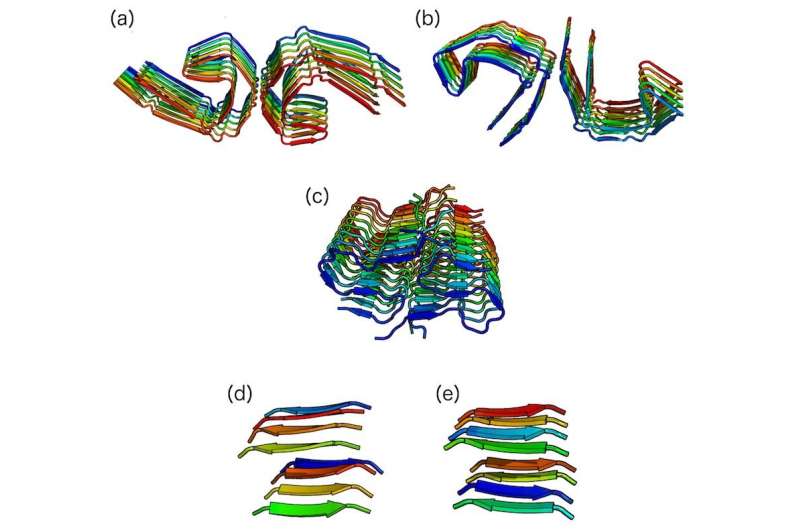
"The sequences of these three amyloids fibrils vary greatly in composition and length, but all participate in an aggregation pathway that results in these fibril structures. It is common to study amyloid proteins individually; however, by studying three very different amyloid fibrils together, we were able to determine stabilizing forces that were common among them. This gives us new directions for understanding amyloids in general and a better understanding of how some amyloids result in disease states," said Brown.
With these new insights, researchers can begin to design drugs to break up the amyloid fibrils or prevent them from forming in the first place.
"Therapeutic intervention will be most helpful if researchers can design drugs that prevent the fibril formation," said Lemkul.
Darcy Davidson, the first author on the paper and a first-year graduate student in Lemkul's lab, began her research on microtubule-associated tau as part of her rotation project.
"Both of my grandfathers were diagnosed with Alzheimer's, so this research is personal and important to me. These discoveries can help researchers develop better drugs to target specific areas of amyloid fibril formation, and this is exciting in terms of future treatment for Alzheimer's," said Davidson.
Davidson is currently continuing her research on microtubule-associated protein tau focusing on protein folding and how single proteins begin to aggregate to form an amyloid fibril.
Lemkul's team is interested in future collaboration with researchers to test these discoveries in the laboratory and in animal models; researchers can then begin to design drugs to target and prevent the formation of amyloid fibrils.
Amyloid formation is associated with a variety of human diseases, including Alzheimer's disease (AD), type 2 diabetes (T2D), Parkinson's, rheumatoid arthritis, Huntington's disease, and more.
More information: Darcy S. Davidson et al. Insights into Stabilizing Forces in Amyloid Fibrils of Differing Sizes from Polarizable Molecular Dynamics Simulations, Journal of Molecular Biology (2018). DOI: 10.1016/j.jmb.2018.05.020