Researchers use RNA to target pancreatic beta cells
Investigators at the University of Miami Miller School of Medicine have designed RNA molecules that home to human pancreatic beta cells, the cells that produce insulin and are destroyed in patients with type 1 and type 2 diabetes. This targeting approach could help researchers study how these diseases progress and provide new ways to deliver treatments. The study was published on April 5 in Nature Communications.
"We created a tool to deliver agents specifically to human beta cells," said associate professor Paolo Serafini, Ph.D., senior author on the paper. "This gives us new capabilities. We can use PET or CT scans to count beta cells and measure disease progression, and we can deliver therapies directly to beta cells without causing side effects in other parts of the body."
Type 1 diabetes is caused by an autoimmune response that targets pancreatic beta cells. Over time, the immune system destroys these critical insulin-producing cells and the body can no longer control glucose levels, leading to severe blood vessel and organ damage. While insulin can be an effective treatment, it is not a cure.
Type 2 diabetes has a different progression. Cells develop insulin resistance, meaning beta cells must work harder to produce more of the hormone. Eventually, beta cells can die of exhaustion, once again dysregulating glucose metabolism.
Researchers studying these diseases have never been able to get a good census on how many beta cells are alive in the pancreas at any given moment. They rely on proxy measures, such as glucose levels, which provide only limited information on how these conditions evolve. Scientists trying to develop treatments are often hamstrung by the toxicity and other side effects associated with systemic therapies—they've wanted a system that can ferry treatments directly to beta cells.
"If we treat patients with non-targeted drugs that encourage beta cells to proliferate, those agents can affect many other tissues and potentially cause cancer," said Dr. Serafini. "Similarly, if we try to protect beta cells from the autoimmune response, non-targeted treatments will cause systemic immunosuppression, leaving patients vulnerable to pathogens and, again, cancer."
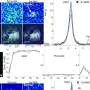
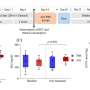
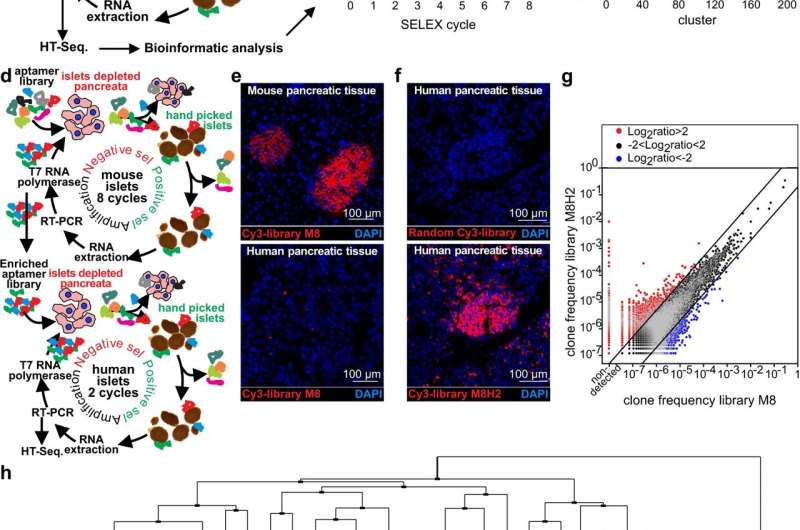
Dr. Serafini and colleagues solved these problems by creating two distinct aptamers, short RNA molecules that selectively target beta cells and ignore other tissues. In turn, scientists can attach additional molecules to illuminate beta cells (for imaging) or to treat the disease.
This will give scientists new opportunities to create targeted treatments that encourage beta cells to proliferate or locally dampen immunity without causing systemic side effects. In addition, the ability to precisely measure beta cell mass will improve future studies.
"It's going to be cheaper and easier to perform clinical trials because we will be able to see exactly what's happening to beta cells in real time," said Dr. Serafini. "In the past, we had to measure glucose or insulin to infer what was happening in the pancreas. Now, we can actually see it."
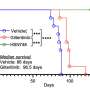
The researchers have already begun testing potential therapeutic options. They are working with WiNK Therapeutics and the Juvenile Diabetes Research Foundation (JDRF) to conduct preclinical studies to hopefully move these approaches into human trials in the next few years.
"We are working with JDRF on follow-up studies to block the autoimmune response and encourage beta cells to proliferate," said Dr. Serafini. "In theory, with sufficient treatment, we can increase beta cell mass to reverse diabetes and also protect beta cells from autoimmunity so people no longer need insulin."
More information: Dimitri Van Simaeys et al, RNA aptamers specific for transmembrane p24 trafficking protein 6 and Clusterin for the targeted delivery of imaging reagents and RNA therapeutics to human β cells, Nature Communications (2022). DOI: 10.1038/s41467-022-29377-3