Researchers show how blood vessels regroup after stroke
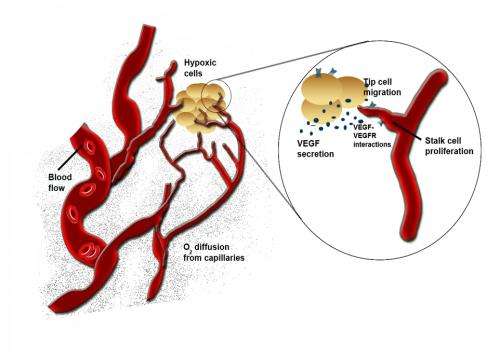
Rice scientists simulate "robot" cells to study the development of microvascular systems in the brain. The goal is to find a way to direct the development of vessels that feed oxygen-starved cells in stroke and neurodegenerative disease patients.
By thinking of cells as programmable robots, researchers at Rice University hope to someday direct how they grow into the tiny blood vessels that feed the brain and help people regain functions lost to stroke and disease.
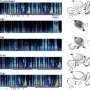
Rice bioengineer Amina Qutub and her colleagues simulate patterns of microvasculature cell growth and compare the results with real networks grown in their lab. Eventually, they want to develop the ability to control the way these networks develop.
The results of a long study are the focus of a new paper in the Journal of Theoretical Biology.
"We want to be able to design particular capillary structures," said Qutub, an assistant professor of bioengineering based at Rice's BioScience Research Collaborative. "In our computer model, the cells are miniature adaptive robots that respond to each other, respond to their environment and pattern into unique structures that parallel what we see in the lab."
When brain cells are deprived of oxygen – a condition called hypoxia that can lead to strokes – they pump out growth factor proteins that signal endothelial cells. Those cells, which line the interior of blood vessels, are prompted to branch off as capillaries in a process called angiogenesis to bring oxygen to starved neurons.
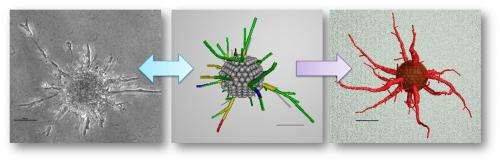
How these new vessels form networks and the shapes they take are of great interest to bioengineers who want to improve blood flow to parts of the brain by regenerating the microvasculature.
"The problem, especially as we age, is that we become less able to grow these blood vessels," Qutub said. "At the same time, we're at higher risk for strokes and neurodegenerative diseases. If we can understand how to guide the vessel structures and help them self-repair, we are a step closer to aiding treatment."
First, they need to understand how individual cells respond to stimuli. To model the process in a computer requires rules, Qutub said. In these simulations, each cell is a "state machine," a unit that goes from one "state" to the next in time based on input. In the case of these endothelial cells, the input comes from vascular endothelial growth factor and/or brain-derived neurotrophic factor, proteins that encourage angiogenesis.
"There's a memory in each of these cells that helps define how they emerge into these very elaborate vasculature structures," she said. The cells that quickly differentiate into tip and stalk cells follow particular rules to advance, grow, divide and branch, depending on input from growth factors and from neighboring cells that can dictate how far and how fast they develop.
For instance, she said, in a chain of endothelial cells, a tip cell at its maximum length can only continue to migrate if the stalk cell immediately behind grows and pushes it forward. Similarly, the team set rules for branching and changes in direction, as well as an "idle" state, all based on observation of real vessel growth.
The researchers modeled a set of endothelial cells growing from a sphere and exposed them to simulated growth factors. They let them grow for what, in real life, would be a period of 24 hours, but in the computer took fractions of a second. They ran tens of thousands of simulations to see how the cells would migrate, proliferate and branch under various conditions.
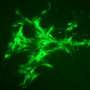
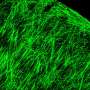
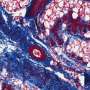
The next step was to figure out which of the computer simulations matched actual behavior. Qutub's lab cultured spheres of human umbilical vein endothelial cells in collagen scaffolds, exposed them to growth factors and took microscope images as they developed networks over several days.
They compared these images with the simulations. The closest matches – of which there are only a few amid thousands of simulations – became the basis for a refined set of rules.
"We know how cells are connected as a function of the growth factors, and there are very distinct patterns to the way these networks are organized," Qutub said. "So when we compare the simulations with the assays, we find parallels that let us classify what we see in the real networks. This gives us a paradigm where we can start to think of actually programming real cells with chemical signaling.
"It opens up a lot of doors," she said. "Now we can think about changing things in the cells or giving them drugs that target pathways and growth factors to induce a particular vasculature structure."
More information: dx.doi.org/10.1016/j.jtbi.2012.11.030