Comprehensive tumour immunity map opens up immunotherapy to more patients
Scientists have developed a new way to map the molecules on tumour cells that flag their presence to the immune system, according to a study published today in eLife.
The findings could make commonly used immunotherapy treatments effective for a much larger population of cancer patients.
Some cancer immunotherapy treatments work by targeting short pieces of proteins called peptides that are displayed in the surface of cancer cells. These peptides are presented on the cell surface by human leukocyte antigens (HLAs) of which there are many different types. However, immunotherapy treatment research tends to focus on a small subset of HLAs only. As not all cancer patients produce these HLAs, they are unable to benefit from existing HLA-based immunotherapies.
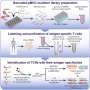
"Most studies have focused on HLA proteins that are commonly found in the general population," explains co-first author Kenji Murata, a postdoctoral fellow at the Princess Margaret Cancer Centre, University Health Network, Toronto, Canada. "We have developed a new technique that allows the sampling of underrepresented HLA proteins to find peptides or antigens that can induce an antitumour immune response. We can then stimulate the patient's own immune cells with those peptides and give them back to the patient to help treat their cancer."
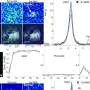
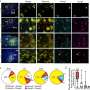
The team began by isolating white blood cells called T cells from eight patients with melanoma. This patient group spanned 25 different types of HLAs, allowing the team to analyse T cell interactions with more than 800 different antigen peptides. All eight patient samples were positive for at least one of the peptide-HLA combinations. The methods developed here also allowed the team to discover new peptides recognised by the T cells.
Next, the team explored whether the T cells that reacted to the peptides could cause an immune response by measuring the production of an immune-activating molecule called interferon. All except two of the antigens stimulated the T cells to produce interferon as well as stimulate the cells to increase in number. These effects were made more robust by adding to the T cells another type of cell—an artificially engineered antigen-presenting cell (APC) bearing the same antigen, which is a common strategy to stimulate T cells.
In the next stage, the team used the artificial APCs to find the exact immunogenic signal that stimulated the T cells from melanoma patients. They found novel peptide fragments related to two different antigens, called MART1 and NY-ESO1, which are known to contain immunological hotspots. They looked at whether they could engineer T cells to target these novel antigens by taking cells that react against the novel antigens and cloning their T cell receptor (TCR) genes. TCRs are cell-surface proteins that allow T cells to react to antigens. "When we added these cloned TCR genes back into newly isolated T cells, we found that the cells were able to recognise and react to the tumour cells," says co-first author Munehide Nakatsugawa, a former postdoctoral fellow at the Princess Margaret Cancer Centre.
"By querying human melanoma-derived T cells and using novel HLA proteins bearing common tumour antigens, we have been able to discover both new and existing immunologically active antigens in tumours," concludes senior author Naoto Hirano, Senior Scientist at the Princess Margaret Cancer Centre. "Our strategy allows for a more complete examination of the immune response and development of novel cancer vaccines and immunotherapies for a broader group of patients not limited by HLA prevalence or tumour mutation burden."
More information: Kenji Murata et al, Landscape mapping of shared antigenic epitopes and their cognate TCRs of tumor-infiltrating T lymphocytes in melanoma, eLife (2020). DOI: 10.7554/eLife.53244