Cancer researchers validate a clinical test for fusion genes
An assay that identifies a peculiar but important abnormality in cancer cells has been developed and validated by researchers at The Ohio State University Comprehensive Cancer Center – Arthur G. James Cancer Hospital and Richard J. Solove Research Institute (OSUCCC – James).
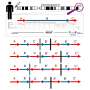
The assay, called OSU-SpARKFuse (Ohio State University-Spanning Actionable RNA Kinase Fusions), detects a genetic change called gene fusions in solid tumors. The assay and its validation are published in the Journal of Molecular Diagnostics.
Gene fusions happen when parts of two different genes join together. Gene fusions can happen, for example, when a piece of one chromosome becomes attached to another. Such chromosome "translocations" can join two genes that together become a major driver of cancer-cell and tumor growth.
Targeted therapies are becoming increasingly available that block the activity of fusion genes, particular those involving kinase genes. Whereas current assays for detecting gene fusions require previous knowledge of both genes involved in the fusion, OSU-SpARKFuse was designed to accurately detect fusions when only one of the genes is known, which allows for the discovery of novel gene fusions.
"We designed OSU-SpARKFuse to meet these needs and to identify patients who are eligible for novel therapies such as FGFR inhibitors or NTRK inhibitors that target gene fusions," says principal investigator Sameek Roychowdhury, MD, PhD, assistant professor in the Division of Medical Oncology at Ohio State.
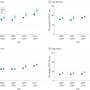
"Along with detecting gene fusions, OSU-SpARKFuse can provide gene-expression analysis, detect single-nucleotide changes and identify alternative splicing events and resistance genes," says first author Julie Reeser, PhD, technical supervisor of the OSUCCC – James Cancer Genomics Laboratory.
"Additionally, OSU-SpARKFuse does not require information regarding the location of the fusion in each gene. It is an accurate, reproducible, cost-effective assay that detects gene fusions across many genes and from the small samples of tumor tissue obtained by biopsy," Reeser adds.
Roychowdhury, Reeser and their colleagues validated the performance of OSU-SpARKFuse using 74 positive and 36 negative control specimens. They included 51 cell line samples, 43 formalin-fixed paraffin-embedded tissues and 16 fresh-frozen tissues. (The assay targets 93 kinase and transcription-factor genes.)
Next, they used the assay to assess gene-fusion status in 95 tissue samples from patients with advanced cancer as part of an Ohio State clinical tumor sequencing study (OSU-13053, NCT02090530) designed and led by Roychowdhury.
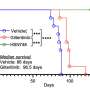
Evaluation of the patient samples revealed a novel fusion of the RET gene (RET- OLFM4) in a patient with small-bowel cancer. It also led to the discovery of a KLK2-FGFR2 fusion in a patient with prostate cancer. The patient was then treated with a fibroblast growth factor receptor inhibitor.
"The use of OSU-SpARKFuse in clinical laboratories will help expand the knowledge base of gene fusions in solid tumors, and it could directly affect patient care by detecting therapeutically actionable targets," Roychowdhury says.
More information: Julie W. Reeser et al. Validation of a Targeted RNA Sequencing Assay for Kinase Fusion Detection in Solid Tumors, The Journal of Molecular Diagnostics (2017). DOI: 10.1016/j.jmoldx.2017.05.006